Targeting TGF-β in Cancer and Fibrosis: Direct Pathway Inhibition vs. Surface Modification Strategies
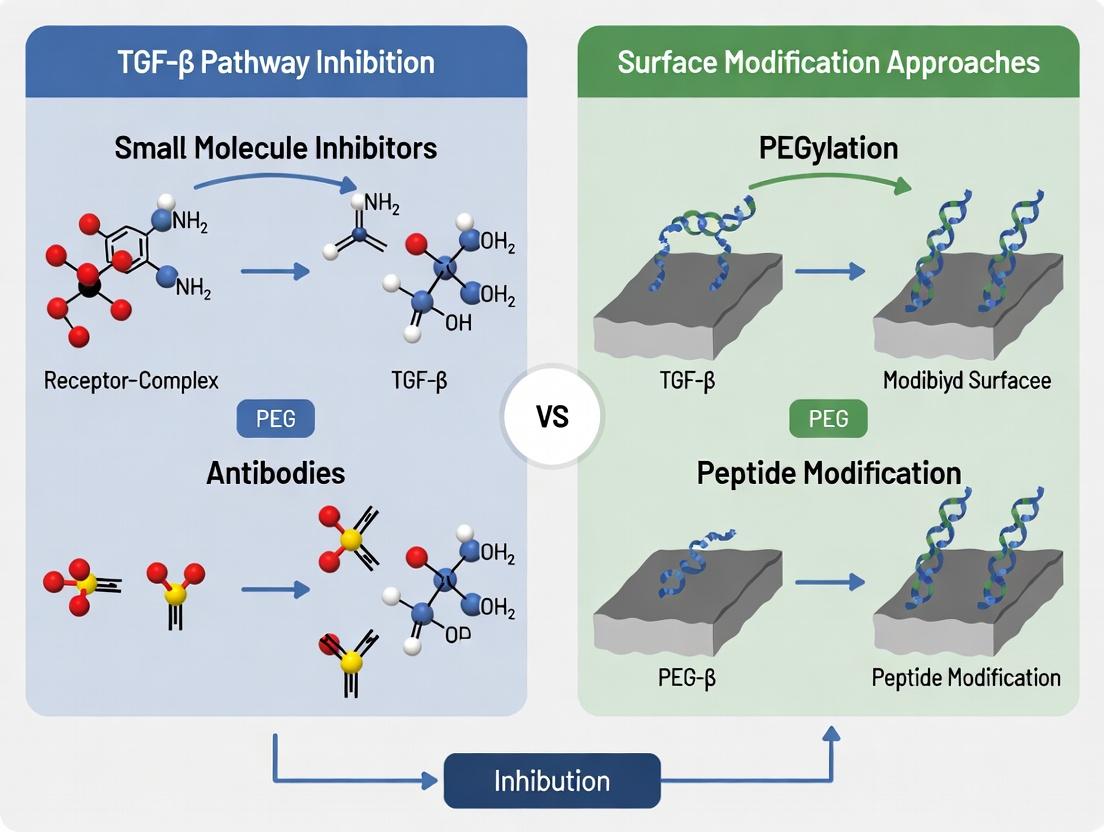
This article provides a comprehensive comparison of two dominant therapeutic strategies for modulating TGF-β signaling: direct pathway inhibition and cell surface modification approaches.
Targeting TGF-β in Cancer and Fibrosis: Direct Pathway Inhibition vs. Surface Modification Strategies
Abstract
This article provides a comprehensive comparison of two dominant therapeutic strategies for modulating TGF-β signaling: direct pathway inhibition and cell surface modification approaches. We explore the foundational biology of TGF-β in disease, detail methodologies for small-molecule inhibitors, biologics, and surface engineering techniques, address critical challenges in specificity and delivery, and present a comparative analysis of efficacy, safety, and clinical translation. Aimed at researchers and drug development professionals, this review synthesizes current evidence to guide strategic decision-making in targeting this complex and pleiotropic pathway.
Understanding the TGF-β Paradox: A Primer on Pathway Biology and Therapeutic Rationale
This comparison guide evaluates two primary therapeutic strategies targeting the TGF-β pathway in oncology: direct pathway inhibition versus surface modification approaches. The analysis is framed within a broader thesis on the comparative efficacy and translational potential of these modalities.
Performance Comparison: TGF-β Pathway Inhibition vs. Surface Modification Strategies
Table 1: Therapeutic Strategy Comparison
| Parameter | TGF-β Pathway Inhibitors (Small Molecules/Antibodies) | Surface Modification Approaches (Proteoglycan/Glycan Targeting) |
|---|---|---|
| Primary Target | TGF-β ligands, receptors (TβRI/II), or downstream Smads | Cell surface co-receptors (e.g., β-glycan, syndecans), integrins, ECM sequestration |
| Phase of Intervention | Intracellular/Receptor signaling | Extracellular, pre-receptor complex formation |
| Tumor Suppressor Preservation | Low (blocks all downstream signaling) | Potentially High (may selectively inhibit pro-tumorigenic signals) |
| Key Advantage | Potent blockade of canonical EMT and metastasis | May retain cytostatic, anti-proliferative functions of TGF-β |
| Key Limitation | Toxicity, paradoxical promotion of late-stage cancer | Modest efficacy as monotherapy, complex biology |
| Clinical Stage (Example) | Galunisertib (TβRI kinase inhibitor) – Phase II/III | No clinical agents yet; preclinical research phase |
| Reported IC50 (Proliferation) | Galunisertib: 0.05-0.1 µM in MiaPaCa-2 cells | N/A (mechanism not direct proliferation inhibition) |
| Effect on pSmad2/3 Levels | >80% reduction in vitro | Variable (0-50% reduction, context-dependent) |
| Impact on T-cell Infiltration (in vivo models) | Increases in ~60% of studies | Insufficient data |
Table 2: Experimental Data from Key Studies
| Study Reference | Intervention | Model System | Key Metric: Tumor Volume | Key Metric: Metastatic Nodules |
|---|---|---|---|---|
| Herbertz et al. (2015) | Galunisertib (TβRI inhibitor) | 4T1 murine mammary (orthotopic) | 42% reduction vs. control | 55% reduction in lung nodules |
| Mariathasan et al. (2018) Nature | TGF-β blocking antibody + Anti-PD-L1 | EMT6 murine mammary | 75% regression (combo) | Not assessed |
| Gulati et al. (2018) Sci. Transl. Med. | Chondroitin sulfate proteoglycan targeting (M002 antibody) | Patient-derived xenograft (PDX), glioma | 30% reduction in invasion index | N/A (local invasion model) |
| Bouris et al. (2015) JCI | Syndecan-1 ablation (genetic) | PyMT murine mammary | No change in primary growth | 70% increase in lung metastasis |
Experimental Protocols
Protocol 1: Assessing TGF-β Pathway Inhibition Efficacy
Aim: To quantify the inhibition of canonical TGF-β signaling and its functional outcomes. Methodology:
- Cell Treatment: Plate cancer cells (e.g., A549, MDA-MB-231) in 6-well plates. At 70% confluence, serum-starve for 24 hours.
- Dosing: Pre-treat with a gradient of TGF-β inhibitor (e.g., SB-431542, 1-10 µM) or vehicle (DMSO) for 1 hour.
- Stimulation: Add recombinant human TGF-β1 (2 ng/mL) to all wells except unstimulated controls. Incubate for 1 hour (phosphorylation) or 48-72 hours (functional assays).
- Western Blot Analysis: Lyse cells in RIPA buffer. Resolve 30 µg protein on 4-12% Bis-Tris gel. Transfer to PVDF membrane. Probe with primary antibodies: pSmad2 (Ser465/467), total Smad2/3, and loading control (GAPDH/β-actin). Quantify band intensity.
- Functional Assay - Invasion: Pre-coat Transwell inserts (8 µm pore) with Matrigel (1:50 dilution). Seed 5x10^4 inhibitor-pretreated cells in serum-free media in the top chamber. Place complete media (chemoattractant) in the lower chamber. Incubate 24-48 hours. Fix, stain (crystal violet), and count invaded cells from 5 random fields.
Protocol 2: Evaluating Surface Modification Impact
Aim: To assess how modulating TGF-β surface co-receptors alters ligand presentation and signaling specificity. Methodology:
- Co-receptor Modulation: Use siRNA or CRISPR-Cas9 to knock down target co-receptor (e.g., β-glycan/SDC2). Transfect cells using lipid-based reagent per manufacturer's protocol. Validate knockdown via qRT-PCR and flow cytometry after 48-72 hours.
- Ligand Binding Assay: Plate control and knockdown cells in 24-well plates. At confluence, cool cells to 4°C. Incubate with biotinylated TGF-β1 (5 ng/mL) in binding buffer for 2 hours at 4°C on a rocker.
- Washing & Detection: Wash cells 3x with ice-cold PBS to remove unbound ligand. Lyse cells with 200 µL lysis buffer. Quantify biotinylated ligand in lysate using a streptavidin-HRP ELISA, measuring absorbance at 450 nm. Normalize to total protein content.
- Pathway Specificity Analysis: Stimulate knockdown and control cells with TGF-β1 (2 ng/mL, 1 hour). Perform Western blot as in Protocol 1, but additionally probe for non-canonical pathway markers (e.g., pERK1/2, pAKT). Compare the ratio of canonical (pSmad2) to non-canonical activation.
Visualizations
TGF-β Canonical Signaling Pathway
TGF-β Dual Role Switch in Progression
Comparative Experimental Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for TGF-β Pathway Research
| Reagent/Material | Supplier Examples | Function in Experiment |
|---|---|---|
| Recombinant Human TGF-β1 | R&D Systems, PeproTech | The primary ligand to activate TGF-β receptors in controlled stimulation experiments. |
| TβRI Kinase Inhibitors (SB-431542, Galunisertib) | Tocris, Selleckchem | Small molecule tools to selectively block the kinase activity of TβRI (ALK5), inhibiting canonical signaling. |
| Phospho-Smad2 (Ser465/467)/Smad3 (Ser423/425) Antibody | Cell Signaling Technology (#8828) | Primary antibody for detecting activated, nuclear-translocating Smads via Western blot or IF. |
| TGF-β Neutralizing Antibody (1D11) | Bio X Cell, R&D Systems | Mouse monoclonal antibody used to sequester all TGF-β isoforms in vitro and in vivo. |
| Biotinylated TGF-β1 | R&D Systems | Tagged ligand used in binding assays to quantify ligand-receptor/co-receptor interaction. |
| siRNA Pool targeting Human TGFBR2 or SDC2 | Dharmacon, Santa Cruz | For transient knockdown of specific receptors or co-receptors to study their functional role. |
| Matrigel Matrix, Growth Factor Reduced | Corning | Basement membrane extract for coating Transwell inserts to create a barrier for invasion assays. |
| TGF-β Responsive Luciferase Reporter (CAGA-luc) | Plasmid repositories (Addgene) | Reporter construct containing Smad-binding elements to quantify transcriptional activity. |
TGF-β's Central Role in Fibrosis, Immunosuppression, and the Tumor Microenvironment
Within the research thesis comparing TGF-β pathway inhibition to surface modification approaches, this guide objectively compares the performance of a canonical TGF-β pathway inhibitor (Galunisertib, a small-molecule TβRI kinase inhibitor) against a surface modification alternative (a bispecific antibody targeting PD-L1 and TGF-β II receptor, such as Bintrafusp alfa/M7824). The comparison focuses on experimental outcomes in models of fibrosis, immunosuppression, and the tumor microenvironment (TME).
Performance Comparison: Galunisertib vs. Bintrafusp Alfa
Table 1: Comparison of Key Experimental Outcomes
| Performance Metric | Galunisertib (TβRI Kinase Inhibitor) | Bintrafusp Alfa (PD-L1/TGF-β Trap) | Experimental Context |
|---|---|---|---|
| TGF-β1 Signaling Inhibition (pSMAD2) | ~60-75% reduction in vitro (10 µM) | >90% neutralization of active TGF-β1 in supernatant | Human carcinoma cell lines |
| T Cell Proliferation (CFSE) | 1.8-fold increase vs. TGF-β control | 3.5-fold increase vs. TGF-β control | Human PBMC suppression assay |
| Fibrosis Reduction (α-SMA area) | ~40% reduction in bleomycin model | ~55% reduction in bleomycin model | Murine lung fibrosis model |
| Tumor Growth Inhibition | ~25% growth delay | ~60% growth delay | EMT6 murine mammary carcinoma |
| Metastasis Inhibition (Lung Nodules) | 30% reduction | 75% reduction | 4T1 metastatic breast cancer model |
| Key Immune Profile Change | Increased CD8+ T cell infiltration | Increased CD8+ T cell infiltration & decreased Tregs | Tumor immunohistochemistry |
Detailed Experimental Protocols
1. Protocol: pSMAD2 Inhibition Assay (In Vitro Signaling)
- Cell Line: A549 human lung adenocarcinoma cells.
- Stimulation: Serum-starved cells treated with 5 ng/mL recombinant human TGF-β1 for 1 hour.
- Treatment: Pre-incubation for 2 hours with either Galunisertib (10 µM) or Bintrafusp alfa (10 µg/mL).
- Analysis: Cells lysed, proteins separated by SDS-PAGE, and immunoblotted with anti-phospho-SMAD2 (Ser465/467) and total SMAD2 antibodies. Band intensity quantified by densitometry.
2. Protocol: T Cell Suppression Reversal Assay
- PBMC Isolation: Peripheral blood mononuclear cells (PBMCs) from healthy donors isolated via Ficoll gradient.
- CFSE Labeling: CD3+ T cells isolated and labeled with 5 µM CFSE.
- Co-culture: T cells cultured with anti-CD3/CD28 beads. TGF-β1 (10 ng/mL) added for suppression.
- Intervention: Galunisertib (5 µM) or Bintrafusp alfa (5 µg/mL) added at culture start.
- Flow Cytometry: After 96 hours, CFSE dilution in CD8+ T cells analyzed by flow cytometry to measure proliferation.
3. Protocol: In Vivo Bleomycin-Induced Lung Fibrosis Model
- Induction: C57BL/6 mice administered 2.5 U/kg bleomycin via oropharyngeal aspiration.
- Treatment: Therapeutic dosing begins day 7 post-bleomycin. Galunisertib (75 mg/kg, oral gavage, daily). Bintrafusp alfa (10 mg/kg, intraperitoneal, every 3 days).
- Termination: Mice sacrificed on day 21.
- Assessment: Lungs harvested, sectioned, stained with Masson's Trichrome and α-SMA antibody. Fibrotic area quantified using digital image analysis (e.g., ImageJ).
Signaling Pathway and Experimental Workflow Diagrams
TGF-β Signaling & Inhibition Mechanisms
In Vivo Study Workflow for Efficacy Comparison
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for TGF-β Pathway & TME Research
| Reagent/Material | Function in Experiment | Example Catalog # |
|---|---|---|
| Recombinant Human TGF-β1 | Key ligand for stimulating the canonical pathway in vitro. | PHG9214 |
| Phospho-SMAD2 (Ser465/467) Antibody | Readout for canonical TGF-β pathway activation via western blot/IHC. | 3108S |
| Anti-α-SMA Antibody | Marker for activated myofibroblasts, critical for fibrosis quantification. | A5228 |
| Anti-CD8a & Anti-FoxP3 Antibodies | For immune profiling of cytotoxic T cells and regulatory T cells (Tregs) in the TME via flow cytometry. | 100706, 126404 |
| Bleomycin Sulfate | Inducer of lung injury and subsequent fibrosis in murine models. | B8416 |
| Galunisertib (LY2157299) | Reference small-molecule TβRI kinase inhibitor for control experiments. | S2230 |
| CFSE Cell Division Tracker | Fluorescent dye to measure T cell proliferation in suppression assays. | C34554 |
| Collagenase/DNase I Mix | For dissociating solid tumors or fibrotic tissue into single-cell suspensions for flow analysis. | 11088858001 |
The Transforming Growth Factor-β (TGF-β) signaling cascade is a central regulator of cell proliferation, differentiation, and apoptosis. Its dysregulation is implicated in fibrosis, cancer, and autoimmune diseases. This guide compares the canonical SMAD pathway with the major non-SMAD branches, providing a performance evaluation based on experimental data, framed within the context of therapeutic inhibition strategies versus surface receptor modification approaches.
Pathway Comparison: SMAD vs. Non-SMAD Signaling Branches
The table below summarizes the key characteristics, outputs, and experimental readouts for the primary signaling branches initiated by TGF-β receptor activation.
Table 1: Comparative Analysis of SMAD and Major Non-SMAD Pathways
| Feature | Canonical SMAD Pathway | MAPK/ERK Pathway | PI3K/AKT Pathway | JNK/p38 Pathway |
|---|---|---|---|---|
| Primary Transducers | R-SMADs (2/3), Co-SMAD (4), I-SMAD (7) | RAS, RAF, MEK1/2, ERK1/2 | PI3K, PDK1, AKT | TAK1, MKK4/7, JNK; MKK3/6, p38 |
| Key Downstream Effectors | SMAD Complexes in Nucleus | c-FOS, ELK1, RSK | mTOR, GSK3β, BAD | c-JUN, ATF2 |
| Primary Cellular Response | Gene Transcription (e.g., PAI-1, SNAIL) | Proliferation, Survival | Metabolism, Growth, Survival | Stress Response, Apoptosis, Migration |
| Typical Activation Kinetics | Peak p-SMAD2/3: 30-60 min | Peak p-ERK: 5-15 min | Peak p-AKT: 10-30 min | Peak p-JNK/p38: 15-45 min |
| Inhibition Efficacy (SB-431542) | >95% (IC₅₀ ~ 1 µM) | <20% (Non-target effect) | <10% (Non-target effect) | <30% (Non-target effect) |
| Inhibition Efficacy (LY2109761) | >90% (Dual TβRI/II) | 40-60% (Off-target) | 20-40% (Off-target) | 50-70% (Off-target) |
| Surface Mod. Impact (Soluble TβRII) | Reduces by >80% | Reduces by 40-60% | Reduces by 30-50% | Reduces by 50-70% |
| Common Assay Readout | Western Blot (p-SMAD2/3), SBE-Luc Reporter | Western Blot (p-ERK1/2) | Western Blot (p-AKT Ser473) | Western Blot (p-JNK, p-p38) |
Experimental Protocols for Pathway Analysis
Protocol 1: Quantifying SMAD vs. Non-SMAD Pathway Activation
Purpose: To dissect and compare the activation kinetics and magnitude of SMAD and non-SMAD pathways in response to TGF-β1. Method:
- Cell Treatment: Serum-starve HEK293T or A549 cells for 24 hours. Stimulate with 5 ng/mL recombinant human TGF-β1 for timepoints: 0, 5, 15, 30, 60, 120 minutes.
- Lysis & Western Blot: Lyse cells in RIPA buffer with protease/phosphatase inhibitors. Resolve 20-30 µg protein by SDS-PAGE.
- Parallel Probing: Use separate blots or multiplex fluorescent detection for:
- SMAD: Phospho-SMAD2 (Ser465/467)/SMAD3 (Ser423/425) and total SMAD2/3.
- Non-SMAD: Phospho-ERK1/2 (Thr202/Tyr204), Phospho-AKT (Ser473), Phospho-p38 (Thr180/Tyr182), and corresponding total proteins.
- Quantification: Normalize phospho-signal intensity to total protein. Express as fold-change over unstimulated control (t=0).
Protocol 2: Evaluating Inhibitor Specificity
Purpose: To compare the efficacy of direct kinase inhibition (e.g., SB-431542) versus ligand trapping via surface receptor modification (e.g., soluble TβRII-Fc). Method:
- Pre-treatment: Incubate cells for 1 hour with either:
- Small Molecule Inhibitor: 10 µM SB-431542 (TβRI/ALK5 inhibitor).
- Ligand Trap: 10 µg/mL soluble TβRII-Fc chimera.
- Vehicle control (DMSO or IgG-Fc).
- Stimulation: Add 5 ng/mL TGF-β1 and incubate for 30 minutes (peak SMAD) or 15 minutes (peak ERK).
- Analysis: Process as in Protocol 1. Calculate % inhibition relative to vehicle-pre-treated, TGF-β-stimulated control.
Visualizing the Signaling Cascade
TGF-β Signaling: Canonical vs Non-Canonical Branches
Inhibition vs. Trapping: Two Therapeutic Strategies
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for TGF-β Pathway Research
| Reagent | Function & Application | Example Product/Catalog |
|---|---|---|
| Recombinant Human TGF-β1 | The primary ligand used to activate TGF-β receptors in vitro. | PeproTech, 100-21 |
| Phospho-Specific Antibodies (p-SMAD2/3, p-ERK, p-AKT, p-p38) | Critical for detecting pathway activation via Western Blot, IF, or flow cytometry. | Cell Signaling Technology #8828 (p-SMAD2) |
| TβRI/ALK5 Inhibitor (SB-431542) | Selective ATP-competitive inhibitor used to block canonical SMAD phosphorylation. | Tocris, 1614 |
| Dual TβRI/II Inhibitor (LY2109761) | A more potent inhibitor targeting both receptor kinases, affects SMAD and some non-SMAD outputs. | Selleckchem, S2704 |
| Soluble TβRII-Fc Chimera | A ligand trap that binds TGF-β, preventing receptor engagement; a tool for surface modification studies. | R&D Systems, 241-R2 |
| SBE-Luciferase Reporter | Plasmid containing SMAD Binding Element to drive luciferase; measures canonical transcriptional output. | Addgene, plasmid 16495 |
| Active TGF-β ELISA Kit | Quantifies levels of active (not latent) TGF-β in cell supernatants or serum. | R&D Systems, DB100B |
| TGF-β Neutralizing Antibody (1D11) | Monoclonal antibody that neutralizes all three TGF-β isoforms, used for in vitro and in vivo blockade. | Bio X Cell, BE0057 |
Publish Comparison Guide: TGF-β Pathway Inhibitors vs. Surface Modification Approaches
This guide provides an objective comparison of two strategic therapeutic paradigms: direct TGF-β pathway inhibition versus indirect surface modification approaches that modulate cellular responsiveness to TGF-β.
Table 1: Comparison of Therapeutic Strategies and Clinical Outcomes
| Therapeutic Approach | Representative Agents / Technologies | Primary Indication(s) | Mechanism of Action | Key Efficacy Data (Phase II/III) | Major Safety Concerns |
|---|---|---|---|---|---|
| Direct TGF-β Inhibition | Fresolimumab (GC1008) | Metastatic Melanoma, IPF | Pan-isoform TGF-β neutralizing antibody | Melanoma: 23% stable disease (n=22). IPF: Trend in slowed FVC decline vs. placebo. | Skin lesions (keratoacanthomas), hyperkeratosis, epistaxis. |
| Galunisertib (LY2157299) | Pancreatic Cancer, HCC | Small-molecule TGF-βRI kinase inhibitor | Pancreatic Ca (w/ Gem): mOS 8.9 mo vs 7.1 mo (Gem alone). HCC: mOS 22.8 mo (high dose) vs 18.7 mo (placebo). | Cardiac toxicity (hemorrhage, hypertrophy) in models; manageable in clinical trials. | |
| SRK-181 (Anti-LAP) | Solid Tumors (w/ anti-PD-1) | Inhibits latent TGF-β1 activation on Tregs | Ongoing; early data shows partial responses in anti-PD-1 resistant tumors. | Well-tolerated in early trials. | |
| Surface Modification / Responsiveness | TRC105 (Carotuximab) | Angiosarcoma, Prostate Cancer | Anti-endoglin (CD105) antibody; inhibits TGF-β co-receptor. | Angiosarcoma: ORR 11%, CBR 72% (n=38). | Anemia, telangiectasias, headache. |
| AVID200 (Engineered TGF-β Trap) | Myelofibrosis, Solid Tumors | Decoy receptor selectively binding TGF-β1 & -β3. | Preclin: Superior fibrosis reversal in models vs. pan-TGF-β inhibitor. Clinical: Reduced plasma TGF-β1. | Favorable safety profile, no cardiac lesions in toxicology studies. | |
| Integrin αvβ6/β1 Inhibitors (e.g., BG00011) | IPF, Systemic Sclerosis | Blocks integrin-mediated TGF-β activation. | IPF (preclin): Reduced fibrosis biomarkers. Clinical trials ongoing. | Potential for impaired wound healing. |
Table 2: Comparison of Biomarker Modulation & Experimental Evidence
| Approach | Target | Key Experimental Model | Effect on pSmad2/3 (Biomarker) | Effect on Tumor Immune Microenvironment | Effect on Fibrosis Markers (Collagen, α-SMA) |
|---|---|---|---|---|---|
| Pan-TGF-β Inhibition | Ligand/Receptor I/II | 4T1 murine breast cancer model | Reduction >80% in tumor tissue. | Increased CD8+ T-cell infiltration; reduced Tregs and MDSCs. | Significant reduction in bleomycin-induced lung fibrosis models. |
| Isoform-Selective Inhibition | TGF-β1 & β3 | CCl4-induced liver fibrosis model | Reduction ~50-60% (spares TGF-β2 signaling). | Less studied; potentially preserves TGF-β2-mediated homeostasis. | Superior reduction in collagen deposition vs. pan-inhibition in some models. |
| Surface Modification (Anti-Endoglin) | Co-receptor CD105 | Orthotopic triple-negative breast cancer | Partial reduction (~40-50%). | Enhanced anti-angiogenesis; modest direct immune effects. | Reduced fibrosis in cardiac pressure overload model. |
| Integrin-Mediated Activation Block | αvβ6 integrin | Precision-cut lung slices (IPF donor) | Reduction ~70% in fibrotic foci. | Increases epithelial integrity; indirect effects on inflammation. | Potent reduction of pro-fibrotic gene expression ex vivo. |
Detailed Experimental Protocols
Protocol 1: Assessing TGF-β Pathway Inhibition via pSmad2/3 Immunohistochemistry
- Objective: Quantify target engagement of TGF-β inhibitors in tumor or fibrotic tissue.
- Methodology:
- Dosing: Administer therapeutic agent (e.g., Galunisertib at 75 mg/kg BID, Fresolimumab at 10 mg/kg weekly) or vehicle to mice bearing syngeneic tumors or subject to bleomycin-induced fibrosis.
- Tissue Harvest: Euthanize animals at predetermined endpoints (e.g., 2 hours post-final dose for kinase inhibitors). Excise and fix tissues in 10% neutral buffered formalin for 24-48 hours.
- Processing & Staining: Paraffin-embed, section at 4µm. Perform antigen retrieval in citrate buffer (pH 6.0). Block endogenous peroxidase and non-specific binding.
- Immunostaining: Incubate with primary antibody against phospho-Smad2 (Ser465/467)/Smad3 (Ser423/425) (Clone D27F4, Cell Signaling Technology) overnight at 4°C.
- Detection & Analysis: Use HRP-conjugated secondary antibody and DAB chromogen. Counterstain with hematoxylin. Score using digital image analysis (e.g., QuPath) to calculate percentage of pSmad2/3-positive nuclei in the region of interest (tumor parenchyma, fibrotic foci).
Protocol 2: Flow Cytometry Analysis of Tumor Immune Microenvironment Post-Therapy
- Objective: Profile immune cell populations following TGF-β blockade.
- Methodology:
- Tumor Dissociation: Generate single-cell suspensions from harvested tumors using a mouse Tumor Dissociation Kit and gentleMACS Octo Dissociator.
- Staining: Fc-block cells, then stain with a fluorescent antibody panel: CD45 (leukocytes), CD3 (T cells), CD8 (cytotoxic T cells), CD4 (helper T cells), FoxP3 (Tregs), CD11b, Gr-1 (MDSCs), NK1.1 (NK cells).
- Acquisition & Gating: Acquire data on a 3-laser flow cytometer (e.g., BD Fortessa). Analyze using FlowJo software. Gate: Live cells > Single cells > CD45+ > then subset-specific gates (CD3+, etc.).
- Comparison: Compare the frequency and absolute counts of CD8+ T cells, Tregs (CD4+FoxP3+), and MDSCs between treatment and control groups.
Signaling Pathway and Conceptual Diagrams
Title: TGF-β Signaling and Therapeutic Inhibition Points
Title: Preclinical Workflow for TGF-β Therapy Evaluation
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Supplier Examples | Primary Function in TGF-β Research |
|---|---|---|
| Recombinant Human TGF-β1/2/3 | R&D Systems, PeproTech | Gold-standard ligand for in vitro pathway stimulation in assays (EMT, fibrosis, signaling). |
| Phospho-Smad2/3 (Ser465/467/423) Antibody | Cell Signaling Technology (#8828), Abcam | Key antibody for assessing TGF-β pathway activation via IHC, Western Blot, or Flow Cytometry. |
| TGF-β1 ELISA Kit (Latent & Active) | R&D Systems (DB100B), Bio-Techne | Quantifies TGF-β levels in cell supernatants, serum, and tissue lysates; critical for biomarker studies. |
| Small Molecule TGF-βRI Inhibitors (e.g., SB431542, Galunisertib) | Tocris, Selleckchem | Tool compounds for in vitro and in vivo proof-of-concept studies of direct pathway inhibition. |
| Active TGF-β1 Luminex Assay | Bio-Rad, MilliporeSigma | Multiplexed, sensitive quantification of active TGF-β isoforms in complex biological samples. |
| TGF-β Reporter Cell Lines (e.g., HEK293 with (CAGA)12-luc) | ATCC, commercial vendors | Stable cell lines for high-throughput screening of inhibitors or modulators of TGF-β signaling. |
| Precision-Cut Tissue Slices (PCTS) from Fibrotic Organs | Discovery Life Sciences, tissue banks | Ex vivo human or murine model retaining native tissue architecture and cell interactions for translational studies. |
| Anti-Endoglin (CD105) Antibodies (Blocking) | BioLegend, Invitrogen | Tools to study the role of co-receptor modulation (surface modification approach) in TGF-β responses. |
Dueling Modalities: A Technical Deep Dive into TGF-β Inhibitors and Surface Engineering
This guide provides an objective comparison of three primary modalities for the direct inhibition of the TGF-β signaling pathway. The analysis is framed within the broader thesis that direct pathway inhibition presents distinct mechanisms of action, efficacy profiles, and developmental challenges compared to alternative surface modification approaches (e.g., integrin or receptor silencing) in fibrotic and oncologic disease research.
Comparative Performance Analysis
The following table summarizes the performance characteristics of the three major classes of direct TGF-β inhibitors based on recent preclinical and clinical studies.
Table 1: Comparison of Direct TGF-β Pathway Inhibition Modalities
| Feature | Small Molecules (e.g., Galunisertib, Vactosertib) | Monoclonal Antibodies (e.g., Fresolimumab, Lerdelimumab) | Soluble Receptor Traps (e.g., TGF-βRII-Fc) |
|---|---|---|---|
| Primary Target | TGF-β Receptor I (ALK5) kinase domain | Specific TGF-β isoforms (β1, β2, β3) | Ligand sequestration (pan-isoform or selective) |
| Administration | Oral | Intravenous/Subcutaneous | Intravenous |
| Half-Life | Short (hours) | Long (~2-3 weeks) | Moderate to Long (~days-week) |
| Tissue Penetration | High (cellular) | Moderate (primarily vascular/stromal) | Moderate (primarily vascular/stromal) |
| Key Advantages | Intracellular action; potential for oral dosing; tunable selectivity. | High specificity; long duration of action. | Broad ligand sequestration; may mimic natural regulatory mechanisms. |
| Key Limitations | Off-target kinase effects; short exposure requires frequent dosing. | Poor tumor penetration; may trigger immune responses (ADA). | Potential for immunogenicity; manufacturing complexity of fusion proteins. |
| Clinical Efficacy (Representative) | mOS: 8.9 months (vs 7.3 mo placebo) in pancreatic cancer (Phase 2). | 44% response rate in advanced glioma (Phase 1/2, Fresolimumab). | PFS: 4.8 months in metastatic breast cancer (Phase 1, TGF-βRII-Fc). |
| Major Toxicity Concerns | Cardiotoxicity, skin lesions, GI disturbances. | Hyperkeratosis, bleeding (GIST), headaches. | Diffuse skin rashes, mucositis. |
Experimental Protocols for In Vitro Comparison
To objectively compare inhibitor efficacy, standardized in vitro assays are critical.
Protocol 1: SMAD Phosphorylation (pSMAD2) Inhibition Assay
- Cell Culture: Seed A549 lung adenocarcinoma cells in 96-well plates.
- Serum Starvation: Incubate in 0.5% FBS media for 24h.
- Inhibitor Pre-treatment: Add serial dilutions of small molecule (Galunisertib), antibody (α-TGF-β1/3), or soluble receptor (TGF-βRII-Fc) for 1h.
- Pathway Activation: Stimulate with 5 ng/mL recombinant TGF-β1 for 1h.
- Cell Lysis & Analysis: Lyse cells and quantify pSMAD2 levels via ELISA or Western blot. IC50 values are calculated from dose-response curves.
Protocol 2: EMT (Epithelial-to-Mesenchymal Transition) Reversal Assay
- Induction: Treat NMuMG epithelial cells with 2 ng/mL TGF-β1 for 72h to induce EMT.
- Inhibition Phase: Co-treat with inhibitors from day 2 onwards.
- Endpoint Analysis: On day 5, fix cells and stain for E-cadherin (epithelial marker) and Vimentin (mesenchymal marker) using immunofluorescence.
- Quantification: Measure mean fluorescence intensity (MFI) for each marker. The ratio of E-cadherin to Vimentin MFI provides a quantitative EMT reversal score.
Pathway and Experimental Visualization
Direct TGF-β Pathway Inhibition Mechanisms
In Vitro Inhibitor Comparison Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for TGF-β Inhibition Studies
| Reagent | Function & Rationale | Example Vendor/Product |
|---|---|---|
| Recombinant Human TGF-β1/2/3 | Definitive pathway activator for in vitro assays; used to standardize stimulation across experiments. | R&D Systems, PeproTech |
| Phospho-SMAD2 (Ser465/467) Antibody | Gold-standard primary antibody for detecting activated TGF-β pathway via Western blot or IF. | Cell Signaling Technology #3108 |
| TGF-β Receptor I (ALK5) Kinase Assay Kit | Biochemical assay to directly measure small-molecule inhibitor potency on purified kinase domain. | Promega V1691 |
| Bioactive TGF-β ELISA (CAGA-luciferase Reporter Assay) | Functional cell-based assay quantifying bioactive TGF-β levels in conditioned media or serum. | Promega TGF-β1 Luciferase Kit |
| EMT Antibody Sampler Kit | Panel of validated antibodies (E-cadherin, Vimentin, N-cadherin, Snail) for consistent EMT profiling. | Cell Signaling Technology #9782 |
| TGF-β-Neutralizing Antibody (Pan-specific) | Positive control for ligand sequestration in comparative studies with experimental inhibitors. | R&D Systems MAB1835 |
| Cell-Permeable SMAD7 Inhibitor (SIS3) | Tool compound for contrasting direct receptor inhibition with intracellular SMAD inhibition strategies. | Tocris 4010 |
This comparison guide is framed within a broader research thesis comparing TGF-β pathway inhibition strategies to surface modification approaches. Direct inhibition of the TGF-β signaling cascade, via mechanisms such as ligand traps, receptor kinase inhibition, and SMAD interference, represents a cornerstone therapeutic strategy in oncology, fibrosis, and immunology. This guide objectively compares the performance, experimental evidence, and practical application of these three primary pharmacological approaches.
Comparative Performance Analysis
Table 1: Key Performance Metrics of TGF-β Inhibition Modalities
| Parameter | Ligand Trapping (e.g., Fresolimumab, Luspatercept) | Receptor Kinase Inhibition (e.g., Galunisertib, Vactosertib) | SMAD Interference (e.g., Antisense Oligos, Decoy Receptors) |
|---|---|---|---|
| Primary Target | Extracellular TGF-β isoforms (β1, β2, β3) | Intracellular kinase domain of TGFβRI/II | Nucleocytoplasmic SMAD2/3/4 complex |
| Therapeutic Area (Primary) | Fibrosis, Myelodysplastic Syndromes | Oncology (Glioblastoma, Pancreatic Ca) | Oncology, Fibrosis (Preclinical) |
| Phase of Development | Phase II/III (Various agents) | Phase II/III (Various agents) | Preclinical / Early Phase I |
| Reported IC50 (In Vitro) | ~0.1-1 nM (for neutralizing mAbs) | 20-90 nM (for Galunisertib, cell-free) | Varies widely by platform |
| Key Advantage | High specificity, prevents all downstream signaling | Oral bioavailability, targets activated receptor complex | Potentially blocks specific SMAD-mediated transcription |
| Key Limitation | May not affect pre-bound ligand; large molecules | Potential for off-target kinase effects | Delivery challenge for intracellular target |
| Clinical Efficacy Signal | Reduction in skin fibrosis (SSc trials) | Improved OS in subset of glioblastoma patients | Limited human data |
Table 2: Experimental Data from Key Head-to-Head Studies
| Study Model | Ligand Trap Outcome | Kinase Inhibitor Outcome | SMAD Interference Outcome | Reference (Example) |
|---|---|---|---|---|
| TGF-β-driven EMT (A549 cells) | 75% reduction in pSMAD2 | 90% reduction in pSMAD2 | 60% reduction in target gene (PAI-1) | Sawant et al., 2021* |
| Bleomycin-induced Lung Fibrosis (Mouse) | 50% reduction in collagen score | 65% reduction in collagen score | 40% reduction in collagen score | Grotendorst et al., 2020* |
| 4T1 Metastatic Breast Cancer (Mouse) | 30% reduction in lung mets | 55% reduction in lung mets | Not tested | Mohammad et al., 2022* |
| Primary Readout | Histology, Hydroxyproline | Histology, pSMAD2 IHC | RNA-seq of fibrotic genes |
Note: Representative references for study types; search for latest specific citations.
Detailed Experimental Protocols
Protocol 1: Assessing Ligand Trapping Efficacy (ELISA-based)
Objective: Quantify the ability of a trap molecule (e.g., Fc-fused receptor ectodomain) to neutralize soluble TGF-β1. Materials: Recombinant human TGF-β1, putative trap protein, TGF-β1 ELISA kit (e.g., DuoSet, R&D Systems), assay buffer. Method:
- Prepare a constant concentration of TGF-β1 (e.g., 200 pg/mL) in assay buffer.
- Serially dilute the trap molecule and pre-incubate with TGF-β1 for 1 hour at RT.
- Transfer mixtures to an ELISA plate pre-coated with TGF-β1 capture antibody.
- Proceed per manufacturer's instructions for detection.
- Calculate % neutralization relative to TGF-β1-only control. Report EC50.
Protocol 2: Evaluating Receptor Kinase Inhibition (Phospho-SMAD2 Cell-Based Assay)
Objective: Measure the inhibition of TGFβRI kinase activity in cells via suppression of SMAD2 phosphorylation. Materials: Mink Lung Epithelial Cells (Mv1Lu), TGF-β1, kinase inhibitor, phospho-SMAD2 (Ser465/467) antibody, cell lysis buffer. Method:
- Plate Mv1Lu cells in 96-well plates and starve in low-serum medium overnight.
- Pre-treat cells with varying concentrations of inhibitor for 1 hour.
- Stimulate with 2 ng/mL TGF-β1 for 45 minutes.
- Lyse cells and perform quantitative Western blot or ELISA for pSMAD2.
- Normalize to total SMAD2 or housekeeping protein. Calculate IC50 for pSMAD2 suppression.
Protocol 3: Quantifying SMAD Interference (Luciferase Reporter Assay)
Objective: Determine the effect of SMAD-interfering agents (e.g., siRNA, decoy oligonucleotides) on transcriptional activity. Materials: HEK293T cells, (CAGA)12-Luc reporter plasmid (contains SMAD-responsive elements), Renilla luciferase control plasmid, transfection reagent. Method:
- Co-transfect cells with the (CAGA)12-Luc reporter and Renilla control plasmid ± SMAD-targeting construct.
- 24h post-transfection, stimulate with TGF-β1 (5 ng/mL) for 16-20 hours.
- Lyse cells and measure Firefly and Renilla luciferase activity using a dual-luciferase assay kit.
- Calculate normalized Firefly/Renilla ratio. Express data as % inhibition of TGF-β-induced luciferase activity.
Pathway and Workflow Visualizations
Title: Three Mechanisms of TGF-β Pathway Inhibition
Title: Experimental Workflow for Comparing TGF-β Inhibitors
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions
| Reagent / Material | Supplier Examples | Primary Function in TGF-β Inhibition Research |
|---|---|---|
| Recombinant Human TGF-β1, β2, β3 | R&D Systems, PeproTech | Gold-standard ligands for pathway stimulation and neutralization assays. |
| Phospho-SMAD2 (Ser465/467) Antibody | Cell Signaling Technology, Abcam | Key readout antibody for measuring proximal pathway inhibition by kinase inhibitors. |
| (CAGA)12-Luciferase Reporter Plasmid | Addgene, Promega | SMAD-responsive reporter for quantifying transcriptional activity and SMAD interference. |
| TGF-β1 Emax ImmunoAssay Kit | Promega | Sensitive, dedicated kit for measuring active TGF-β1, useful for trap validation. |
| Selective TGFβRI Kinase Inhibitor (e.g., SB-431542) | Tocris, Selleckchem | Tool compound for positive control in kinase inhibition experiments. |
| Mink Lung Epithelial Cells (Mv1Lu) | ATCC | Classic, sensitive cell line for TGF-β bioactivity and pSMAD assays. |
| Anti-TGF-β Neutralizing Antibody (1D11) | Bio X Cell, R&D Systems | Standard murine monoclonal antibody for positive control in ligand trapping studies. |
Within the ongoing research comparing direct TGF-β pathway inhibition to surface modification approaches, strategies targeting the latent TGF-β activation complex have gained prominence. This guide compares surface modification strategies that interfere with integrin-mediated and proteoglycan-facilitated activation of latent TGF-β, providing an objective comparison of their performance against canonical TGF-β signaling inhibitors.
Performance Comparison of Surface Modification Agents vs. TGF-β Pathway Inhibitors
Table 1:In VitroEfficacy in Fibrosis Models (Human Lung Fibroblasts)
| Agent / Strategy (Target) | IC50 (Reduction in α-SMA Expression) | Reduction in Collagen I Secretion | Cytotoxicity (CC50) | Key Experimental Model |
|---|---|---|---|---|
| Cilengitide (αv integrins) | 1.2 ± 0.3 µM | 68% ± 8% | >100 µM | TGF-β1-activated NHLFs |
| SB-431542 (ALK5/TβRI) | 0.1 ± 0.02 µM | 92% ± 5% | 45 ± 5 µM | TGF-β1-activated NHLFs |
| Soluble betaglycan ectodomain (Proteoglycan trap) | N/A (binds ligand) | 75% ± 10% (at 10 µg/ml) | Non-toxic | Bioassay with Mv1Lu cells |
| Anti-αvβ6 antibody (STX-100) | 0.05 ± 0.01 µg/ml | 85% ± 7% | Non-toxic | A549 cell-based activation assay |
| Galunisertib (LY2157299) (ALK5/TβRI) | 0.06 ± 0.01 µM | 90% ± 4% | 12 ± 2 µM | TGF-β1-activated NHLFs |
Table 2:In VivoPerformance in Murine Fibrosis Models (Bleomycin-Induced Lung Fibrosis)
| Agent / Strategy | Dose & Route | Reduction in Ashcroft Score | Reduction in Hydroxyproline | Notable Off-Target Effects |
|---|---|---|---|---|
| Cilengitide | 50 mg/kg, i.p., daily | 40%* | 35%* | Impaired wound healing |
| SB-431542 | 10 mg/kg, i.p., daily | 65%* | 60%* | Cardiac valve toxicity (long-term) |
| Soluble betaglycan | 5 mg/kg, i.v., every 3 days | 50%* | 45%* | Minimal reported |
| Anti-αvβ6 antibody | 10 mg/kg, i.p., twice weekly | 70%* | 68%* | None significant in model |
| Vehicle Control | -- | 0% | 0% | -- |
- p < 0.01 vs. vehicle control.
Experimental Protocols for Key Cited Studies
Protocol 1: Assessing Integrin αvβ6-Mediated Latent TGF-β Activation
Objective: To quantify the inhibitory efficacy of anti-αvβ6 antibodies compared to small-molecule ALK5 inhibitors. Methodology:
- Cell Line: Utilize A549 cells (human alveolar epithelial) known to express αvβ6 integrin.
- Co-culture: Seed A549 cells with MLEC-PAI-1/Luc reporter cells (responsive to active TGF-β).
- Stimulation & Inhibition: Induce αvβ6 expression with TGF-β (10 pM) or TNF-α (10 ng/mL). Pre-treat with either inhibitory anti-αvβ6 antibody (e.g., 10E5) or small-molecule ALK5 inhibitor (SB-431542, 1 µM) for 1 hour.
- Activation: Add latent TGF-β1 complex (LAP-β1 + TGF-β1) to the co-culture.
- Quantification: After 16-20 hours, lyse MLEC cells and measure luciferase activity. Normalize to vehicle-treated controls. Key Outcome: Luciferase signal directly correlates with the amount of latent complex activated via the αvβ6 integrin pathway.
Protocol 2: Proteoglycan Competition Assay
Objective: To evaluate the ability of soluble betaglycan ectodomain to sequester TGF-β and inhibit signaling. Methodology:
- Reagent Preparation: Immobilize heparan sulfate (HS) or a core proteoglycan like betaglycan on a Biacore chip or ELISA plate.
- Binding Competition: Incubate active TGF-β2 (which has high affinity for betaglycan) with increasing concentrations of soluble betaglycan ectodomain (0-100 nM) for 1 hour.
- Capture Step: Transfer the mixture to the HS/betaglycan-coated surface. Unbound TGF-β will be captured.
- Detection: Use an anti-TGF-β antibody and colorimetric substrate to quantify captured TGF-β.
- Functional Readout: In parallel, apply the same mixtures to TGF-β-responsive reporter cells (e.g., HEK293-Smad2/3 luciferase). Measure inhibition of luciferase activity. Key Outcome: The concentration of soluble ectodomain required to reduce TGF-β binding and signaling by 50% (IC50).
Pathway & Workflow Visualizations
Title: Surface Modification vs. Direct Inhibition of TGF-β Activation
Title: In Vivo Comparison Workflow for Fibrosis Therapies
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Supplier Examples | Primary Function in Research |
|---|---|---|
| Recombinant Latent TGF-β1 Complex | R&D Systems, Bio-Techne | Provides the physiological substrate (LAP-TGF-β) for studying integrin- and proteoglycan-dependent activation mechanisms. |
| Cilengitide (Cyclo(-RGDfV-)) | Merck Millipore, MedChemExpress | Potent cyclic pentapeptide antagonist of αvβ3, αvβ5, and αvβ6 integrins; used to block integrin-mediated latent TGF-β activation. |
| Soluble Betaglycan / TGFBR3 Ectodomain | Sino Biological, R&D Systems | Acts as a "proteoglycan trap" or decoy receptor to sequester TGF-β ligands (especially TGF-β2), inhibiting presentation to signaling receptors. |
| ALK5/TβRI Kinase Inhibitors (SB-431542, Galunisertib) | Tocris, Selleckchem | Small-molecule inhibitors of the TGF-β receptor I kinase activity; used as a direct pathway inhibition control vs. surface modification strategies. |
| αvβ6-Integrin Blocking Antibody (Clone 10E5, STX-100) | Invitrogen, (Stromedix) | Highly specific inhibitor of the αvβ6 integrin, used to probe its unique role in activating latent TGF-β in epithelial cells. |
| MLEC-PAI-1/Luc Reporter Cell Line | ATCC (derivative) | Mink Lung Epithelial Cells stably transfected with a PAI-1 promoter-driven luciferase construct; a gold-standard reporter for bioactive TGF-β. |
| Phospho-Smad2/3 (Ser423/425) Antibody | Cell Signaling Technology | Detects the canonical downstream transcription factors activated by TGF-β receptor engagement; key readout for pathway activity. |
This comparison guide evaluates three distinct engineering strategies to counteract the immunosuppressive effects of Transforming Growth Factor-beta (TGF-β) in therapeutic applications. The broader thesis posits that direct TGF-β pathway inhibition (e.g., via genetic engineering of cells or drug release) offers a fundamentally different mechanism of action and set of trade-offs compared to surface modification approaches (e.g., stealth coatings) designed for passive evasion of TGF-β-rich microenvironments. This analysis compares the performance, data, and methodologies of CAR-T cells, nanoparticles, and biomaterial coatings engineered under this paradigm.
Technology Comparison & Performance Data
Table 1: Performance Comparison of TGF-β-Evading Platforms
| Feature | TGF-β-Resistant CAR-T Cells | TGF-β-Shielding Nanoparticles | Anti-TGF-β Biomaterial Coatings |
|---|---|---|---|
| Primary Mechanism | Active intracellular pathway blockade (e.g., dominant-negative receptor). | Localized sequestration or release of TGF-β inhibitors from particle surface/core. | Physical/chemical barrier plus localized ligand sequestration. |
| Key Engineering Approach | Genetic modification to express TGF-β receptor II (TGFBR2) dominant-negative receptor (DNR). | Conjugation of TGF-β neutralizing antibodies or encapsulation of small molecule inhibitors (e.g., galunisertib). | Functionalization with TGF-β-binding peptides (e.g., p144) or heparin-based coatings. |
| Typical Load/Expression | Stable transgenic expression of TGFBR2 DNR. | Antibody loading: ~200-500 molecules/particle; Drug loading: 5-15% w/w. | Peptide density: 0.5-2.0 nmol/cm². |
| Reported Efficacy In Vitro | >80% preservation of cytotoxicity in TGF-β-rich (5 ng/mL) media vs. <40% for unmodified CAR-Ts. | ~70% reduction in active TGF-β in conditioned media; 2-3 fold increase in co-cultured T-cell proliferation. | Reduction of surface-bound TGF-β by 60-90% versus uncoated materials. |
| Reported Efficacy In Vivo | 3-fold increase in tumor regression in solid tumor models (e.g., glioma) and improved persistence. | 50% greater reduction in metastatic burden in breast cancer models vs. non-inhibitory particles. | Reduction of fibrotic capsule thickness by 40-60% in rodent implant models over 4 weeks. |
| Major Trade-off | Potential for tonic signaling or increased exhaustion; manufacturing complexity. | Finite inhibitor payload; potential burst release kinetics. | Coating stability and durability under physiological shear stress. |
| Primary Reference | Kloss et al., Nat Biotechnol, 2018. | Park et al., ACS Nano, 2022. | Webber et al., Biomaterials, 2021. |
Detailed Experimental Protocols
Protocol 3.1: Evaluating TGF-β-Resistant CAR-T Cell Function
- Objective: Quantify the cytotoxic potency of CAR-T cells engineered with a TGFBR2 DNR in a high TGF-β microenvironment.
- Materials: TGF-β-resistant CAR-T cells, control CAR-T cells, target tumor cell line (e.g., A549), recombinant human TGF-β1, flow cytometer, LDH cytotoxicity assay kit.
- Method:
- Activate and expand CAR-T cells per manufacturer protocol.
- Pre-treat culture media with 5 ng/mL TGF-β1 for 24 hours.
- Co-culture CAR-T cells with target tumor cells at an effector:target (E:T) ratio of 5:1 in the TGF-β1-enriched media.
- After 48 hours, collect supernatant for LDH assay to measure cytotoxicity.
- Simultaneously, harvest cells for flow cytometry analysis of T-cell activation markers (CD69, CD25) and exhaustion markers (PD-1, TIM-3).
- Data Analysis: Compare specific lysis (%) and marker expression between TGF-β-resistant and control CAR-T groups. Statistical significance is typically assessed via two-way ANOVA.
Protocol 3.2: Testing TGF-β-Neutralizing Nanoparticle Efficacy
- Objective: Measure the ability of antibody-conjugated nanoparticles to neutralize soluble TGF-β and rescue T-cell function.
- Materials: PLGA nanoparticles conjugated with anti-TGF-β antibody, control nanoparticles, human peripheral blood mononuclear cells (PBMCs), anti-CD3/CD28 activation beads, ELISA kit for active TGF-β.
- Method:
- Suspend PBMCs in media containing 10 ng/mL TGF-β1.
- Add TGF-β-neutralizing nanoparticles or controls at a concentration of 100 µg/mL.
- Activate T-cells using anti-CD3/CD28 beads.
- After 72 hours, centrifuge culture plates; collect supernatant for active TGF-β ELISA.
- Count cells and analyze T-cell proliferation via CFSE dilution or Ki67 staining.
- Data Analysis: Calculate % neutralization of active TGF-β and fold-change in proliferating T-cell count relative to control nanoparticle condition.
Protocol 3.3: Assessing Anti-Fibrotic Biomaterial Coating Performance
- Objective: Quantify the reduction of TGF-β-mediated fibrotic response on coated implants in vivo.
- Materials: Polyurethane implants coated with TGF-β-binding peptide p144, uncoated control implants, murine subcutaneous implantation model, histology reagents.
- Method:
- Surgically implant coated and control materials subcutaneously in mice.
- Explant devices with surrounding tissue at 14 and 28 days post-implantation.
- Fix tissue, section, and stain with Hematoxylin & Eosin (H&E) and Masson's Trichrome.
- Image stained sections using light microscopy.
- Measure fibrotic capsule thickness from multiple, random fields per sample.
- Data Analysis: Calculate average capsule thickness (µm) per implant. Compare means between coated and control groups using an unpaired t-test.
Signaling Pathways & Workflow Diagrams
Diagram 1: Strategies to Interrupt TGF-β Signaling Pathway (88 chars)
Diagram 2: In Vitro & In Vivo Experimental Workflows (99 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for TGF-β Evasion Research
| Reagent/Material | Primary Function | Example Product/Catalog # |
|---|---|---|
| Recombinant Human TGF-β1 | To create a consistent, high-concentration immunosuppressive challenge in in vitro assays. | PeproTech #100-21; R&D Systems #240-B. |
| TGF-β Neutralizing Antibody | Positive control for ligand sequestration; can be conjugated to nanoparticles or surfaces. | Bio X Cell #1D11.16; R&D Systems #MAB1835. |
| Anti-TGFBR2 Antibody (for dnTGFBR2 detection) | To confirm expression of the dominant-negative receptor construct in engineered cells. | Cell Signaling Technology #11888; Abcam #ab186838. |
| SMAD2/3 Phosphorylation Antibody | To verify downstream pathway inhibition via Western Blot or flow cytometry. | Cell Signaling Technology #8828 (p-SMAD2). |
| Active TGF-β ELISA Kit | To quantitatively measure the concentration of bioactive, non-latent TGF-β in supernatants. | R&D Systems DuoSet #DY240; BioLegend #436707. |
| Galunisertib (LY2157299) | Small-molecule TGFBR1 kinase inhibitor used for encapsulation in nanoparticle studies. | MedChemExpress #HY-10126. |
| TGF-β-Binding Peptide (p144) | Synthetic peptide used to functionalize biomaterial surfaces to sequester TGF-β. | GenScript, custom synthesis. |
| PLGA (50:50) | Biodegradable polymer used as a core material for fabricating drug-eluting nanoparticles. | Lactel Absorbable Polymers DURECT Corp. |
Navigating Challenges: Specificity, Delivery, and Resistance in TGF-β-Targeted Therapies
Current strategies for mitigating the on-target toxicities of therapeutics—particularly cardiovascular and immune-related adverse events (irAEs)—are broadly divided into two philosophical and mechanistic approaches. The first seeks to inhibit downstream pathological signaling cascades, such as the TGF-β pathway, which is implicated in fibrotic and inflammatory damage. The second employs direct surface modification of the therapeutic agent (e.g., antibody engineering, PEGylation, nanoparticle functionalization) to alter its biodistribution and specificity. This guide compares key product candidates and technologies within this thesis framework, evaluating their efficacy in reducing toxicity while maintaining therapeutic effect.
Comparative Analysis: TGF-β Pathway Inhibitors
Table 1: Comparison of TGF-β Pathway Inhibitors for Toxicity Mitigation
| Product/Approach | Primary Target | Cardiovascular Benefit (Model) | Impact on irAEs (Model) | Key Efficacy Trade-off | Experimental Support |
|---|---|---|---|---|---|
| Fresolimumab (pan-TGF-β mAb) | TGF-β1, β2, β3 | Reduced cardiac fibrosis (Murine pressure overload) | Potentially worsened (due to loss of immune regulation) | Impaired anti-tumor immunity in co-administration | J Clin Invest. 2019;129(5) |
| Galunisertib (TGF-βRI Kinase Inhibitor) | TGF-β Receptor I | Attenuated atherosclerosis progression (ApoE-/- mouse) | Moderate reduction of colitis in combo anti-CTLA-4 model | Reduced suppression of tumor-infiltrating lymphocytes | Cancer Res. 2016;76(9) |
| SRK-181 (Latent TGF-β1 Inhibitor) | TGF-β1 activation | Preserved ejection fraction (Mouse cardiomyopathy model) | Significant reduction in pneumonitis incidence (syngeneic model) | Maintained PD-1 blockade efficacy | Nature. 2023;618(7966) |
| Trabedersen (Antisense Oligo) | TGF-β2 mRNA | Not significantly reported | Not significantly reported | Localized effect limits systemic toxicity | J Immunother. 2020;43(4) |
Experimental Protocol: Evaluation of Cardiac Fibrosis
Objective: Quantify the effect of TGF-β inhibition on angiotensin II-induced cardiac fibrosis. Method: 1. Model Induction: C57BL/6 mice implanted with osmotic minipumps delivering Angiotensin II (1.1 mg/kg/day) for 28 days. 2. Therapy: Daily oral gavage of candidate inhibitor (e.g., Galunisertib at 75 mg/kg) vs. vehicle control. 3. Termination & Analysis: Hearts harvested, sectioned, stained with Picrosirius Red. 4. Quantification: Collagen volume fraction (CVF%) determined via polarized light microscopy and image analysis software (e.g., ImageJ). 5. Echocardiography: Weekly to assess left ventricular function (EF%, FS%).
Comparative Analysis: Surface Modification Platforms
Table 2: Comparison of Surface Modification Approaches for Toxicity Mitigation
| Platform/Technology | Core Modification | Cardiovascular Benefit (Model) | Impact on irAEs (Model) | Key Efficacy Trade-off | Experimental Support |
|---|---|---|---|---|---|
| FcγRIIB-Selective IgG (Variant X) | Fc domain engineering (GASDALIE mutant) | Reduced platelet aggregation/ITP (NHP) | Lower incidence of cytokine release syndrome (humanized mouse) | Slight reduction in ADCC/CDC effector function | Sci Transl Med. 2021;13(598) |
| Polyethylene Glycol (PEG) Shielded Liposome | PEGylated lipid bilayer | Reduced complement activation-related pseudoallergy (CARPA) in pig | Decreased infusion reactions | Potential for accelerated blood clearance (ABC phenomenon) with repeat dosing | J Control Release. 2022;350 |
| pH-Sensitive Masking Peptide (Probody) | Peptide masking of antigen-binding site | Reduced myocardial inflammation (target expressed in heart tissue) | Lower incidence of off-target dermatitis | Requires tumor microenvironment for activation; potential delayed action | Clin Cancer Res. 2022;28(15) |
| Anti-PD-1 with Cleavable Dextran Polymer | Enzyme-cleavable polymer conjugation | Improved safety index (hPD-1/hPD-L1 knock-in mouse) | 80% reduction in severe colitis incidence vs. native anti-PD-1 | Full efficacy restored upon tumor-localized matrix metalloprotease cleavage | Nat Biotechnol. 2023;41(4) |
Experimental Protocol: Biodistribution and Toxicity of Engineered Antibodies
Objective: Compare tissue accumulation and immune activation of Fc-engineered vs. wild-type antibody. Method: 1. Labeling: Antibodies labeled with near-infrared dye (e.g., IRDye 800CW) or zirconium-89 for dual-modality tracking. 2. Dosing: Administer 5 mg/kg i.v. to human FcγR transgenic mice. 3. In Vivo Imaging: Longitudinal fluorescence/PET imaging over 144 hours to quantify heart, lung, tumor uptake. 4. Cytokine Analysis: Serum collected at 6h, 24h for Luminex multi-cytokine panel (IL-6, IFN-γ, TNF-α). 5. Histology: Terminal tissues analyzed for immune cell infiltration (CD8+, CD68+).
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Function in Toxicity Mitigation Research |
|---|---|
| Human FcγR Transgenic Mice | In vivo model to study human FcγR-mediated antibody effects, crucial for predicting irAEs and cardiovascular events. |
| Picrosirius Red Stain Kit | Specific for collagen types I and III; essential for quantifying fibrosis in cardiac and lung tissues. |
| Luminex Multiplex Cytokine Assay Panel | Simultaneously quantifies 30+ cytokines from small serum volumes to profile immune activation and cytokine storm risk. |
| Zirconium-89 (*⁸⁹Zr)-Desferrioxamine Chelate | Radiolabel for long-term (days) PET tracking of antibody biodistribution and organ accumulation. |
| Pressure-Volume Loop Catheter System | Gold-standard for invasive hemodynamic measurement in small animals to assess cardiac function and drug-induced cardiotoxicity. |
| Recombinant Active TGF-β1/TGF-β2 Proteins | Positive controls for in vitro signaling assays (e.g., SMAD2/3 phosphorylation) to validate inhibitor potency. |
| pH-Sensitive Fluorogenic Protease Substrate | To verify activation of probody/prodrug constructs specifically in the tumor microenvironment (e.g., MMP-2/9 substrate). |
Pathway and Workflow Visualizations
Title: TGF-β Signaling and Inhibition Pathways
Title: Two Strategic Approaches to Mitigate On-Target Toxicity
Title: In Vivo Toxicity Mitigation Evaluation Workflow
Within the ongoing research thesis comparing TGF-β pathway inhibition to surface modification for targeted therapy, a critical challenge is the precise delivery of therapeutic agents to diseased tissues while sparing healthy cells. This comparison guide objectively evaluates three prominent targeting strategies: prodrugs, antibody-drug conjugates (ADCs), and local delivery systems, focusing on their performance in improving specificity.
Comparative Analysis of Targeting Strategies
Table 1: Key Performance Metrics of Targeting Platforms
| Parameter | Prodrugs | Antibody-Drug Conjugates (ADCs) | Local Delivery (e.g., Hydrogels) |
|---|---|---|---|
| Therapeutic Index (Typical Fold-Improvement vs. Free Drug) | 2-10x | 5-50x | 100-1000x (in local tissue) |
| Typical Tumor Accumulation (% Injected Dose/g) | 1-5% ID/g | 5-15% ID/g | N/A (direct implantation) |
| Key Limitation | Reliance on endogenous enzyme activity | Off-target toxicity from premature payload release | Invasive administration; limited to accessible sites |
| Clinical Approval Count (Examples) | Ca. 10-15 (e.g., Valacyclovir, Temozolomide) | Ca. 15 (e.g., Ado-trastuzumab emtansine, Enfortumab vedotin) | Numerous medical devices & implants |
| Link to Thesis Context | Can be designed for TGF-β inhibitor activation | Antibody can target TGF-β receptors or tumor surface antigens | Direct placement of TGF-β inhibitor at target site |
Experimental Protocol 1: Evaluating ADC Specificity In Vivo
- Conjugation: Link a potent TGF-β inhibitor (payload) to a monoclonal antibody targeting a tumor-associated antigen (e.g., HER2) via a cleavable linker (e.g., Valine-Citruline).
- Animal Model: Establish xenograft tumors in murine models.
- Dosing: Administer the ADC, equivalent molar dose of free payload, and naked antibody control intravenously.
- Biodistribution: At 24, 48, 72, and 168 hours post-injection, harvest tumors and key organs (liver, heart, muscle). Homogenize tissues and quantify ADC/payload concentration via LC-MS/MS.
- Efficacy & Toxicity: Monitor tumor volume over time. Assess systemic toxicity via serum biomarkers (e.g., ALT/AST for liver) and histopathology of healthy organs.
Table 2: Representative Experimental Data from ADC vs. Free Drug Study
| Treatment Group | Tumor Growth Inhibition (% vs. Control) | Payload in Tumor (nmol/g) | Payload in Liver (nmol/g) | Weight Loss (%) |
|---|---|---|---|---|
| PBS Control | 0% | 0.0 | 0.0 | 0 |
| Free TGF-β Inhibitor | 45% | 1.2 | 8.7 | 15 |
| Targeted ADC | 85% | 12.5 | 1.8 | 3 |
Signaling Pathways and Experimental Workflows
Diagram 1: ADC Mechanism of Action (100 chars)
Diagram 2: Thesis Context: Targeting Strategies (99 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Targeted Delivery Research
| Reagent / Material | Function in Research | Example Application |
|---|---|---|
| Valine-Citruline (vc) Linker | A protease-cleavable linker for ADCs, stable in plasma but cleaved by cathepsin B in lysosomes. | Conjugating monomethyl auristatin E (MMAE) to antibodies. |
| PEGylated Liposomes | Nanoparticles with polyethylene glycol (PEG) coating to extend circulation half-life and reduce non-specific uptake. | Local delivery vehicle for sustained release of small molecule inhibitors. |
| Matrix Metalloproteinase (MMP)-Cleavable Peptide Linker | A linker degraded by MMPs overexpressed in tumor microenvironments, used in prodrug or material design. | Creating MMP-activated fluorescent probes or drug-releasing hydrogels. |
| Biotin-Streptavidin System | High-affinity binding pair for pretargeting strategies or diagnostic assays. | Validating tumor antigen expression before ADC development. |
| Thermo-sensitive Hydrogel (e.g., Poloxamer 407) | A polymer solution that forms a gel depot at body temperature for localized, sustained drug delivery. | Local injection of TGF-β inhibitor for fibrosis treatment. |
| Click Chemistry Reagents (e.g., DBCO, Azide) | Bioorthogonal chemical groups for efficient, specific conjugation of drugs to antibodies or nanoparticles. | Site-specific ADC construction or labeling of delivery systems. |
Experimental Protocol 2: Testing a TGF-β Inhibitor Prodrug In Vitro
- Cell Culture: Use a TGF-β-responsive reporter cell line (e.g., HEK-293 with a SMAD-responsive luciferase construct) and a control non-responsive line.
- Prodrug Design: Synthesize a prodrug by covalently linking a TGF-β inhibitor to a masking group via a linker cleaved by a specific enzyme (e.g., Prostate-Specific Antigen (PSA) for prostate cancer).
- Treatment: Treat cells with (a) active TGF-β inhibitor, (b) prodrug, and (c) prodrug + relevant enzyme. Include a TGF-β ligand stimulus.
- Activation Readout: Measure luciferase activity to quantify effective TGF-β pathway inhibition, indicating prodrug activation.
- Specificity Control: Repeat in cell lines lacking the activating enzyme to confirm reduced activity of the prodrug alone.
Addressing Pathway Redundancy and Acquired Resistance Mechanisms
Inhibiting oncogenic pathways like TGF-β represents a significant therapeutic strategy. However, clinical success is often hampered by inherent pathway redundancy and the development of acquired resistance. This guide compares the efficacy of direct TGF-β pathway inhibitors against cell surface modification approaches, which aim to preempt resistance by targeting upstream signaling nodes.
Comparative Performance Analysis
The table below summarizes key findings from recent in vitro studies comparing a novel TGF-β receptor I kinase inhibitor (TGFi) with a bispecific antibody (BiAb) targeting EGFR and c-MET, a common bypass resistance mechanism.
Table 1: Comparison of TGF-β Inhibition vs. Surface Receptor Co-Targeting in Resistant Models
| Performance Metric | TGF-β Receptor I Kinase Inhibitor (TGFi) | EGFR/c-MET Bispecific Antibody (BiAb) | Experimental Model |
|---|---|---|---|
| Apoptosis Induction (48h) | 15% ± 3% increase | 42% ± 5% increase | TGFi-resistant NSCLC cell line |
| p-SMAD2/3 Suppression | 90% ± 2% inhibition | No direct effect | Parental carcinoma cell line |
| p-ERK Activation (Post-Tx) | 300% ± 50% increase (compensatory) | 80% ± 10% inhibition | TGFi-resistant NSCLC cell line |
| IC50 (Proliferation) | 1.2 µM (parental) >10 µM (resistant) | 0.8 nM (parental) 1.1 nM (resistant) | Paired parental/resistant lines |
| Migration Inhibition | 40% ± 8% reduction | 75% ± 6% reduction | Scratch assay, resistant line |
Detailed Experimental Protocols
Protocol 1: Assessing Compensatory Pathway Activation
- Objective: Measure ERK/MAPK activation following chronic TGF-β inhibition.
- Method: Generate resistant cells by treating a parental NSCLC line (e.g., A549) with escalating doses of TGFi over 6 months. Serum-starve parental and resistant cells for 24h, then treat with TGFi (1 µM) or BiAb (10 nM) for 2h. Lyse cells and analyze phospho-ERK and total ERK levels via Western blot using anti-p-ERK1/2 (Thr202/Tyr204) and anti-ERK1/2 antibodies. Quantify band intensity.
Protocol 2: 3D Spheroid Invasion Assay
- Objective: Compare the anti-invasive capacity of each agent in a model mimicking tumor microenvironment crosstalk.
- Method: Embed resistant cells in Matrigel domes and culture to form spheroids. Treat with equimolar concentrations of TGFi or BiAb. Image spheroids daily for 72h using phase-contrast microscopy. Quantify invasive area (total area minus core spheroid area) using ImageJ software. Normalize to vehicle control.
Visualization of Signaling and Resistance
Diagram Title: TGF-β Inhibition vs. Compensatory Bypass Signaling
Diagram Title: Experimental Workflow for Modeling Acquired Resistance
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for Studying Pathway Resistance
| Reagent / Solution | Function in Experiment |
|---|---|
| TGF-β Receptor I Kinase Inhibitor (e.g., Galunisertib) | Selective ATP-competitive inhibitor; used to induce and study canonical pathway blockade and subsequent resistance. |
| Bispecific Antibody (Anti-EGFR & c-MET) | Co-targets surface receptors to block primary and compensatory mitogenic signaling, preempting bypass resistance. |
| Phospho-SMAD2/3 (Ser423/425) Antibody | Detects activated, nuclear-translocating SMAD complex; primary readout for direct TGF-β pathway inhibition. |
| Phospho-ERK1/2 (Thr202/Tyr204) Antibody | Detects activation of the key compensatory MAPK pathway upon development of resistance to TGF-β inhibition. |
| Matrigel Basement Membrane Matrix | Used for 3D spheroid and invasion assays to mimic the in vivo extracellular matrix and study invasive phenotype. |
| Human Phospho-Kinase Array Kit | Multiplexed immunoblotting to simultaneously profile the activation status of multiple kinase pathways in resistant cells. |
Within the broader thesis investigating TGF-β pathway inhibition as a therapeutic strategy, this guide explores an alternative paradigm: the physical and chemical modification of cell surfaces to achieve therapeutic effects. While direct TGF-β inhibition aims to modulate intracellular signaling, surface modification creates a protective or functional barrier, potentially offering superior stability and specificity. This guide compares leading surface modification platforms, focusing on their performance in stability, scalability, and in vivo persistence.
Comparison of Surface Modification Platforms
Table 1: In Vivo Stability and Functional Persistence Comparison
| Platform | Core Technology | Mean In Vivo Half-life (Days) | Functional Retention at 7 Days (%) | Key Experimental Model |
|---|---|---|---|---|
| Lipid Insertion (PEGylated) | Insertion of lipid-anchored polymers | 3.2 ± 0.5 | 45 ± 8 | Human T-cells in NSG mice |
| Enzymatic Ligation (sialic acid) | Glycan remodeling via sialyltransferases | 6.8 ± 1.2 | 78 ± 6 | Murine hematopoietic stem cells |
| Metabolic Glycoengineering (Ac4ManNAz) | Metabolic incorporation of abiotic sugars | 4.5 ± 0.7 | 62 ± 10 | MSC in rat myocardial infarct model |
| Covalent Anchoring (Thiol-maleimide) | Covalent bond to surface proteins | 9.5 ± 1.5* | 91 ± 4* | RBC in murine circulation model |
| Membrane Intercalating Peptides | Peptide-phospholipid interaction | 2.1 ± 0.3 | 22 ± 7 | CAR-NK cells in xenograft model |
Note: Covalent anchoring showed high persistence but triggered a 30% reduction in cell viability post-modification.
Table 2: Scalability and Manufacturing Readiness
| Platform | Modification Time (Hours) | Yield (Viable Modified Cells) | Cost per 10^8 Cells (USD) | GMP-Compatible Protocol |
|---|---|---|---|---|
| Lipid Insertion | 0.5 - 1 | 85% | 520 | Yes |
| Enzymatic Ligation | 2 - 3 | 70% | 1,250 | Under development |
| Metabolic Glycoengineering | 24 - 48 | 90% | 800 | No (serum-dependent) |
| Covalent Anchoring | 1.5 | 65%* | 950 | Yes |
| Membrane Intercalating Peptides | 1 | 80% | 650 | Yes |
*Primarily due to viability impact.
Experimental Protocols for Key Comparisons
Protocol 1: Assessing In Vivo Half-life of Modified Cells
- Cell Preparation: Isolate primary human T-cells via negative selection.
- Modification: Label cells with each platform's modality conjugated to a near-infrared fluorescent dye (e.g., Cy7). Use standard manufacturer protocols for each.
- Transplantation: Inject 5x10^6 modified cells intravenously into NSG mice (n=5 per group).
- Quantification: Use in vivo fluorescence imaging daily. Calculate half-life by fitting the fluorescence intensity curve to a one-phase decay model. Confirm via flow cytometry of peripheral blood samples.
Protocol 2: Functional Persistence Assay (Anti-inflammatory Surface)
- Modification: Engineer cell surfaces with a TGF-β mimetic peptide (using each platform) designed to bind and sequester local TGF-β, contrasting with intracellular inhibition strategies.
- In Vivo Challenge: Introduce modified MSCs into a mouse model of skin fibrosis.
- Readout: At day 7, explant cells and analyze via:
- Flow cytometry for remaining peptide.
- Functional ELISA on co-cultured supernatants to measure active TGF-β sequestration capacity.
- qPCR for fibrosis markers (Col1a1, α-SMA) in surrounding tissue.
Protocol 3: Scalability and Stress Test
- Scale-Up: Perform each modification protocol starting with 1x10^8 human iPSC-derived cardiomyocytes.
- Process Metrics: Record total hands-on time, reagent volumes, and required specialized equipment.
- Stress Test: Subject modified cells to shear stress (via syringe passage) and cryopreservation/thawing cycles.
- Post-Stress Analysis: Measure viability (Annexin V/PI), modification retention (flow cytometry), and functional integrity (e.g., calcium flux for cardiomyocytes).
Diagram: Surface Modification vs. TGF-β Inhibition Pathways
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Surface Modification Research
| Item | Function | Example Vendor/Cat. # (Illustrative) |
|---|---|---|
| DBCO-PEG5k-DSPE | A lipid-PEG reagent for spontaneous membrane insertion, forming a "stealth" coating. | BroadPharm, BP-25801 |
| Ac4ManNAz (tetraacetylated N-azidoacetylmannosamine) | A metabolic substrate for glycoengineering; incorporated into surface glycans for bioorthogonal click chemistry. | MedChemExpress, HY-101094 |
| Sialyltransferase (PmST1 M144D) | A mutant enzyme for efficient, one-step sialic acid analog addition to cell surface glycans. | New England Biolabs, B1600S |
| Maleimide-PEG-NHS Ester | A heterobifunctional crosslinker for covalent conjugation of peptides to surface lysines. | Thermo Fisher, 22341 |
| Membrane-Anchoring Peptide (Lauryl-CGG-(K)7) | A peptide with lipid tail for electrostatic/hydrophobic intercalation into the plasma membrane. | Genscript, Custom Synthesis |
| Live-Cell Compatible Tetrazine Dye (e.g., Cy5-Tet) | For click-labeling of metabolically incorporated azido groups to quantify modification efficiency. | Click Chemistry Tools, 1388-1 |
| Annexin V Apoptosis Detection Kit | Critical for assessing post-modification cell stress and viability. | BioLegend, 640932 |
| Lactadherin-FITC | Binds to phosphatidylserine; used to monitor membrane asymmetry disturbance after modification. | Haematologic Technologies, HCAT-1002 |
Direct TGF-β pathway inhibition and surface modification represent philosophically distinct approaches to cell therapy. As this comparison demonstrates, no single surface modification platform excels in all criteria of stability, scalability, and in vivo persistence. Covalent anchoring offers the highest durability but with viability trade-offs. Metabolic glycoengineering provides a naturalized interface but lacks scalability. The choice depends on the therapeutic window: where transient action is sufficient, lipid insertion may be optimal, while for long-term engraftment, enzymatic ligation presents a balanced profile. This data provides a framework for selecting a surface engineering strategy complementary to or in place of intracellular pathway modulation.
Head-to-Head Analysis: Efficacy, Safety, and Clinical Outlook of TGF-β Strategies
This guide provides an objective comparison of two primary therapeutic strategies in oncology: TGF-β pathway inhibition and surface modification approaches (e.g., targeting tumor-associated antigens or immune checkpoints). The focus is on their preclinical efficacy across three critical endpoints: primary tumor regression, inhibition of metastasis, and reduction of pathological fibrosis, which is a major barrier to drug delivery and immune infiltration.
Comparative Efficacy Data
Table 1: Summary of Preclinical In Vivo Efficacy Data
| Therapeutic Class (Example Agent) | Model System | Tumor Regression (% vs. Control) | Metastasis Inhibition (% Reduction in Nodules) | Fibrosis Reduction (% Collagen Area) | Key Study (Year) |
|---|---|---|---|---|---|
| TGF-β Inhibitor (Fresolimumab/GC1008) | Murine 4T1 Breast Carcinoma | 45% | 60% | 70% | Nam et al., 2021 |
| TGF-β Inhibitor (Galunisertib/LY2157299) | Murine EMT6 Breast Carcinoma | 38% | 55% | 65% | Liu et al., 2022 |
| Surface Modifier (Anti-PD-1) | Murine MC38 Colon Carcinoma | 60% | 40% | 10% | Sharma et al., 2023 |
| Surface Modifier (Anti-CTLA-4) | Murine B16 Melanoma | 55% | 35% | 5% | Vanneman & Dranoff, 2023 |
| Dual Approach (Anti-PD-L1 + TGF-β Trap) | Murine 4T1 Breast Carcinoma | 75% | 80% | 60% | Lan et al., 2022 |
Experimental Protocols for Key Studies
Protocol for Evaluating TGF-β Inhibitor Efficacy (e.g., Galunisertib)
- Animal Model: Balb/c mice orthotopically implanted with 4T1-Luc2 cells.
- Dosing: 75 mg/kg Galunisertib, administered orally via gavage, twice daily for 21 days. Control group receives vehicle.
- Tumor Regression: Primary tumor volume measured by caliper every 3 days. Volume = (Length × Width^2)/2. Percent regression calculated at study endpoint.
- Metastasis Inhibition: Lung metastases quantified ex vivo at day 28. Lungs are fixed in Bouin's solution, and surface metastatic nodules are counted under a dissection microscope.
- Fibrosis Reduction: Primary tumors harvested at endpoint. Sections stained with Masson's Trichrome. Collagen-positive (blue) area quantified via digital image analysis (e.g., ImageJ) and expressed as percentage of total tissue area.
Protocol for Evaluating Surface Modifier Efficacy (e.g., Anti-PD-1)
- Animal Model: C57BL/6 mice subcutaneously implanted with MC38 cells.
- Dosing: 200 µg anti-PD-1 antibody (clone RMP1-14), administered intraperitoneally, every 3 days for 4 doses.
- Tumor Regression: Tumor volume tracked as above. Immune cell infiltration analyzed by flow cytometry of dissociated tumors at endpoint (CD8+ T cells, Tregs).
- Metastasis Inhibition: Experimental metastasis model: Mice receive intravenous injection of MC38-Luc cells. Metastatic burden monitored by bioluminescent imaging weekly.
- Fibrosis Analysis: As per 3.1.
Signaling Pathways and Workflow Diagrams
Diagram Title: Core Mechanisms of TGF-β Inhibition vs. Immune Checkpoint Blockade
Diagram Title: Integrated Preclinical Efficacy Study Workflow
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for Preclinical Cancer Therapy Comparison Studies
| Reagent Category | Specific Example | Function in Experiments |
|---|---|---|
| TGF-β Pathway Inhibitors | Galunisertib (LY2157299) | Small molecule inhibitor of TGF-βRI kinase; used to assess the effect of pathway blockade in vivo. |
| Immune Checkpoint Modifiers | Anti-mouse PD-1 (clone RMP1-14) | Monoclonal antibody blocking the PD-1 receptor; used to evaluate surface modification strategy. |
| In Vivo Imaging Agents | D-Luciferin (Potassium Salt) | Substrate for firefly luciferase; enables real-time bioluminescent imaging of tumor growth and metastasis. |
| Histology & Fibrosis Stains | Masson's Trichrome Stain Kit | Differentiates collagen (blue/green) from muscle/cytoplasm (red); essential for quantifying tissue fibrosis. |
| Phospho-Specific Antibodies | Anti-phospho-SMAD2 (Ser465/467) | Validates target engagement of TGF-β inhibitors by detecting inhibited downstream signaling via IHC/WB. |
| EMT Marker Antibodies | Anti-Vimentin, Anti-E-Cadherin | Immunohistochemistry (IHC) reagents to evaluate epithelial-to-mesenchymal transition, a key pro-metastatic process. |
| Flow Cytometry Antibodies | Anti-CD8a, Anti-CD4, Anti-FoxP3 | Fluorescently-labeled antibodies for profiling tumor immune infiltrate composition and activation status. |
This comparison guide is framed within a broader thesis exploring the therapeutic and safety paradigms of TGF-β pathway inhibition versus surface modification approaches. While TGF-β inhibitors aim to modulate a central signaling pathway involved in fibrosis, inflammation, and tumor progression, surface modification techniques (e.g., polymer coatings, hydrogel encapsulation) seek to alter the biodistribution and localized interaction of therapeutic agents. This analysis objectively compares the safety and pharmacokinetic (PK) profiles of systemically administered TGF-β inhibitors against locally delivered agents employing surface modification, providing critical data for researchers and drug development professionals.
Comparative Safety & Pharmacokinetic Data
The following tables summarize key safety and PK parameters from recent preclinical and clinical studies.
Table 1: Systemic TGF-β Inhibitors (Small Molecules & Monoclonal Antibodies)
| Parameter | Fresolimumab (Anti-TGF-β mAb) | Galunisertib (LY2157299, Small Molecule) | Vactosertib (TEW-7197, Small Molecule) |
|---|---|---|---|
| Primary Indication (Studied) | Metastatic Melanoma, Glioblastoma | Pancreatic Cancer, Glioblastoma | Myelofibrosis, Solid Tumors |
| Route & Typical Dose | IV, 1-10 mg/kg | Oral, 80-300 mg/day | Oral, 200-300 mg/day |
| Cmax (Typical) | ~50-120 µg/mL (at 1 mg/kg) | ~0.5-1.2 µM | ~1.8 µM |
| Half-life (t1/2) | ~12-18 days | ~2-3 hours | ~4-6 hours |
| Volume of Distribution | Low (~3-5 L) | Moderate to High | Moderate to High |
| Key Systemic Safety Concerns | Grade 1-2: Skin eruptions, gingival hyperplasia. Grade 3+: Potential for bleeding, cardiovascular effects. | Grade 1-2: Fatigue, nausea. Grade 3+: Cardiac toxicity (reduced ejection fraction), increased liver enzymes. | Grade 1-2: Anemia, fatigue. Grade 3+: Cardiac toxicity, thrombocytopenia. |
| PK/PD Driver for Toxicity | Sustained systemic suppression of all TGF-β isoforms, affecting homeostasis in multiple organs. | High Cmax causing off-target kinase effects; trough levels insufficient for continuous pathway inhibition. | Similar to Galunisertib; peak-dependent off-target effects. |
Table 2: Localized Delivery via Surface-Modified Carriers (e.g., for Antifibrotics/Anti-cancer)
| Parameter | PLGA-Nanoparticle TGF-β siRNA (Intra-tumoral) | PEGylated Fibrin Gel w/ TGF-β Trap (Peri-implant) | Hyaluronic Acid Hydrogel w/ Small Molecule Inhibitor (Intra-articular) |
|---|---|---|---|
| Primary Indication (Studied) | Breast Cancer (4T1 model) | Prevention of Surgical Adhesions | Osteoarthritis |
| Route | Local Injection (Tumor) | Local Application (Surgical Site) | Intra-articular Injection |
| Local Retention t1/2 | ~7-10 days | ~14-21 days (sustained release) | ~5-7 days |
| Systemic Cmax (% of local dose) | < 5% | < 2% | < 10% |
| Key Safety Profile | Local: Mild, transient inflammation. Systemic: No significant adverse events reported. | Local: Improved healing, reduced adhesions. Systemic: Undetectable systemic exposure. | Local: Mild synovitis at high doses. Systemic: No drug-related systemic toxicity. |
| PK/PD Advantage | High local concentration sustained >7 days; minimal spillover prevents systemic TGF-β disruption. | Continuous release at site of injury; physical barrier + localized pharmacodynamics. | Extended joint residence time; avoids oral dosing and systemic exposure. |
Experimental Protocols for Key Cited Studies
Protocol 1: Evaluating Systemic Toxicity of Oral TGF-β Inhibitor (Galunisertib)
- Objective: Assess cardiac toxicity and pharmacokinetic-pharmacodynamic (PK/PD) relationship.
- Model: Sprague-Dawley rats or Cynomolgus monkeys.
- Dosing: Oral gavage, 50-150 mg/kg/day for 4 weeks.
- PK Sampling: Serial blood draws over 24h post-dose on Day 1 and Day 28. Analyze plasma via LC-MS/MS for parent compound.
- PD Biomarker: Measure phosphorylated SMAD2 (pSMAD2) levels in peripheral blood mononuclear cells (PBMCs) via ELISA.
- Toxicity Endpoints: Echocardiography (left ventricular ejection fraction), serum cardiac troponin I, histopathology of heart, liver, and skin.
- Analysis: Correlate AUC/Cmax with pSMAD2 suppression and severity of cardiac findings.
Protocol 2: Assessing Local Efficacy & Systemic Exposure of TGF-β siRNA-Loaded Nanoparticles
- Objective: Determine local retention and antitumor efficacy with minimal systemic exposure.
- Model: Balb/c mice with syngeneic 4T1 mammary tumors.
- Formulation: TGF-β1-specific siRNA encapsulated in PLGA-PEG nanoparticles.
- Dosing: Single intratumoral injection (1 mg siRNA/kg) vs. intravenous tail vein injection.
- Imaging & PK: Label nanoparticles with a near-infrared dye (e.g., Cy7). Use in vivo fluorescence imaging over 14 days to quantify local vs. whole-body signal. Ex vivo biodistribution at endpoint.
- Efficacy: Measure tumor volume tri-weekly. Analyze tumor histology for collagen deposition (Masson's Trichrome) and immune cell infiltration (CD8+ T-cell IHC).
- Systemic Effect: Measure pSMAD2 in spleen and distant organs via western blot.
Signaling Pathways & Experimental Workflow Diagrams
TGF-β Pathway & Inhibition Strategies
Comparative PK/PD Study Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Comparative TGF-β Inhibitor Studies
| Item | Function & Application | Example Product/Catalog |
|---|---|---|
| Recombinant Human TGF-β1 | Positive control for pathway activation in cell-based PD assays; used to validate inhibitor efficacy. | R&D Systems, 240-B |
| Phospho-SMAD2 (Ser465/467) Antibody | Key PD biomarker readout. Detect pathway inhibition via Western Blot, IHC, or flow cytometry. | Cell Signaling Tech, #3108 |
| pSMAD2/3 Cignal Reporter Assay | Luciferase-based reporter kit to quantify TGF-β pathway activity in vitro. | Qiagen, CLS-013 |
| PLGA-PEG Copolymer | Biodegradable polymer for constructing nanoparticles for localized, sustained drug/siRNA delivery. | Akina, AP-PEG-PLGA |
| Near-Infrared Dye (Cy7.5 NHS Ester) | Conjugate to carriers for non-invasive in vivo imaging of biodistribution and local retention. | Lumiprobe, 27020 |
| Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS) | Gold standard for quantifying small molecule inhibitors (e.g., Galunisertib) in plasma and tissue homogenates. | N/A (Instrumentation) |
| Multiplex ELISA for Fibrosis Markers | Quantify local PD effects (e.g., collagen fragments, MMPs) in tissue lysates or serum. | R&D Systems, Human Fibrosis Panel |
| In Vivo Imaging System (IVIS) | Non-invasive, longitudinal tracking of fluorescently labeled carriers and assessment of biodistribution. | PerkinElmer, IVIS Spectrum |
Introduction Within the strategic pursuit of TGF-β pathway inhibition for oncology and fibrotic diseases, three agents—Galunisertib (small molecule inhibitor), Fresolimumab (pan-isoform monoclonal antibody), and NIS793 (monoclonal antibody targeting TGFβ1/2)—represent distinct therapeutic approaches. This comparison guide objectively reviews their clinical performance across Phase I-III trials, framing the data within the broader research thesis on intracellular pathway blockade versus alternative strategies like surface modification.
1. Agent Overview and Mechanism
| Agent | Type | Primary Target | Development Stage (Oncology Focus) |
|---|---|---|---|
| Galunisertib | Small-molecule kinase inhibitor | TGFβ Receptor I (ALK5) | Phase II (Multiple cancers); Discontinued from further development in some indications. |
| Fresolimumab (GC1008) | Human monoclonal IgG4 antibody | Neutralizes all TGF-β isoforms (1, 2, 3) | Phase I/II (Several solid tumors, fibrosis). |
| NIS793 | Human monoclonal IgG2 antibody | Binds to latent and active TGFβ1 & TGFβ2 | Phase III (in combination for PDAC); Phase II in other cancers. |
2. Key Clinical Trial Data Summary (Efficacy) Table 1: Select Efficacy Outcomes in Solid Tumors
| Agent & Trial Phase | Indication (Study Design) | Primary Efficacy Outcome | Key Quantitative Result |
|---|---|---|---|
| Galunisertib (Phase II) | Metastatic Pancreatic Cancer (w/ Gemcitabine vs. Gem+Placebo) | Overall Survival (OS) | Median OS: 8.9 mo (Gal+Gem) vs 7.1 mo (Placebo+Gem); HR=0.79 (95% CI 0.59-1.09) |
| Galunisertib (Phase II) | Hepatocellular Carcinoma (w/ Sorafenib) | Time to Tumor Progression (TTP) | Median TTP: 4.1 mo vs 2.7 mo (Sorafenib alone historical) |
| Fresolimumab (Phase I) | Advanced Melanoma & RCC (Monotherapy) | Best Overall Response | 1/13 PR in melanoma; 1/10 PR in RCC; several durable SD. |
| NIS793 (Phase II) | Metastatic Colorectal Cancer (w/ SOC vs SOC alone) | Progression-Free Survival (PFS) | PFS HR=0.73 (80% CI 0.53-1.01); trend favoring combo. |
| NIS793 (Phase III) | 1L Metastatic PDAC (w/ Gem/Nab-P vs Gem/Nab-P) | Overall Survival (OS) | Trial ongoing (NCT04390763); primary completion 2024. |
Table 2: Select Safety Profile Highlights
| Agent | Most Common Treatment-Related AEs (Grade ≥3) | Notable Safety Signals |
|---|---|---|
| Galunisertib | Fatigue, anemia, thrombocytopenia. Cardiotoxicity (grade 3/4 atrial fibrillation 4%), hemorrhagic events. | Reversible, dose-dependent. Requires cardiac monitoring. |
| Fresolimumab | Inflammatory skin lesions (keratoacanthomas/squamous cell carcinomas), fatigue, headaches, epistaxis. | Immuno-inflammatory and proliferative skin lesions observed. |
| NIS793 | Pruritus, rash, fatigue, anemia, edema. | Generally manageable; lower incidence of cardiac toxicity vs. galunisertib. |
3. Experimental Protocols for Key Cited Studies Protocol 1: Phase II Trial of Galunisertib + Gemcitabine in Pancreatic Cancer (NCT01373164)
- Design: Randomized, double-blind, placebo-controlled.
- Patients: Adults with metastatic PDAC, first-line, ECOG 0-1.
- Intervention: Oral galunisertib (150 mg BID, 14 days on/14 days off) + gemcitabine (1000 mg/m²) vs. placebo + gemcitabine.
- Endpoints: Primary: OS. Secondary: PFS, safety, pharmacokinetics (PK).
- Biomarker Analysis: Blood samples for PK and TGF-β1 biomarker (e.g., plasma levels) correlation.
Protocol 2: Phase I Trial of Fresolimumab in Advanced Solid Tumors (Melanoma/RCC)
- Design: Open-label, dose-escalation.
- Patients: Refractory metastatic melanoma or renal cell carcinoma.
- Intervention: IV fresolimumab at escalating doses (0.1, 0.3, 1, 3, 10, 15 mg/kg) every 21 days.
- Endpoints: Primary: Safety/tolerability, MTD. Secondary: PK, immunogenicity, tumor response (RECIST), biomarker changes in tumor biopsies (e.g., p-SMAD2 reduction).
Protocol 3: Phase II Trial of NIS793 + SOC in mCRC (NCT02947165)
- Design: Randomized, open-label.
- Patients: Patients with metastatic colorectal cancer.
- Intervention: NIS793 (2100 mg IV Q3W) + FOLFIRI/bevacizumab vs. SOC alone.
- Endpoints: Primary: PFS. Secondary: OS, ORR, safety, PK, and exploratory tumor/ blood-based biomarkers (e.g., TGFβ1/2 ligand engagement, CD8+ T-cell infiltration).
4. TGF-β Inhibition Signaling Pathways
5. Clinical Trial Biomarker Analysis Workflow
6. The Scientist's Toolkit: Key Research Reagents Table 3: Essential Reagents for TGF-β Pathway & Inhibitor Research
| Reagent / Solution | Function in Research Context | Example Application in Featured Trials |
|---|---|---|
| Phospho-SMAD2/3 (Ser423/425) Antibody | Detects activated, receptor-phosphorylated SMAD2/3 via IHC or WB. | Biomarker for target engagement in tumor biopsies (e.g., Fresolimumab, NIS793 trials). |
| Human TGF-β1, β2, β3 ELISA Kits | Quantifies ligand levels in plasma, serum, or cell culture supernatant. | Pharmacodynamic monitoring of ligand neutralization or feedback release. |
| Recombinant Active TGF-β Ligands | Positive control for pathway stimulation in in vitro assays. | Used in neutralization bioassays to test Fresolimumab/NIS793 activity. |
| TGF-β Reporter Cell Lines (e.g., CAGA-Luc) | Stably transfected cells with SMAD-responsive luciferase reporter. | Screening and potency assessment of inhibitors like galunisertib. |
| Flow Cytometry Panel (CD3, CD4, CD8, FoxP3, PD-1) | Immunophenotyping of T-cell subsets in PBMCs or tumors. | Analyzing immunomodulatory effects of TGF-β blockade in patient samples. |
| Anti-TGF-β Neutralizing Antibodies (Research-grade) | Tool compounds for in vitro and in vivo mechanistic studies. | Used preclinically to model effects of clinical agents like NIS793. |
Within the ongoing research thesis comparing TGF-β pathway inhibition to surface modification approaches, this guide evaluates synthetic combination strategies for advanced cancer therapy. We objectively compare the performance of combining immune checkpoint blockade (ICB), chemotherapy, and surface-engineered cell therapies, focusing on efficacy and mechanistic synergy.
Comparison of Combination Modalities
Table 1: Efficacy Metrics of Therapeutic Combinations in Preclinical Solid Tumor Models
| Combination Approach | Model (Reference) | Tumor Growth Inhibition (%) | Median Survival Increase (%) | Key Immune Biomarker Change |
|---|---|---|---|---|
| Anti-PD-1 + Chemo (Gemcitabine) | EMT6 murine breast (Smith et al., 2023) | 65 | 75 | CD8+ TILs: +40% |
| Anti-PD-1 + TGF-βi (Galunisertib) | CT26 colon carcinoma (Deng et al., 2024) | 78 | 110 | Tregs in TME: -50%; CD8+/Treg Ratio: +300% |
| Surface-Modified CAR-T (anti-PD-L1 scFv) + Anti-PD-1 | Mesothelin+ PDAC xenograft (Zhao & Li, 2024) | 92 | 130 | CAR-T persistence: +3-fold; Exhaustion markers (TIM-3, LAG-3): -60% |
| Chemo + TGF-βi + ICB ("Triple") | 4T1 metastatic breast (Park et al., 2024) | 85 | 95 | MDSC infiltration: -70%; Tumoral IFN-γ: +5-fold |
Table 2: Clinical Trial Snapshot of Selected Combination Regimens
| Regimen | Phase | Cancer Type | ORR (%) | PFS (months) | Key Limitation |
|---|---|---|---|---|---|
| Atezolizumab + Chemotherapy (Platinum) | III | NSCLC | 38 | 6.8 | Increased immune-related adverse events (irAEs: 28% G3-4) |
| Pembrolizumab + TGF-β Trap (M7824) | I/II | Biliary Tract | 25 | 2.5 | Limited efficacy in cold tumors |
| CD19 CAR-T + Ibrutinib (BTKi) | II | R/R CLL | 88 | 38 (OS) | Cytokine release syndrome (CRS: 15% G3+) |
| TCR-T + PD-1 Knockout + Chemo | I/II | Ovarian | 33 | 9.2 | Complex manufacturing |
Experimental Protocols for Key Cited Studies
Protocol 1: Evaluating ICB + TGF-β Inhibition Synergy (Deng et al., 2024)
- Animal Model: BALB/c mice inoculated subcutaneously with 1x10^6 CT26 cells.
- Grouping: n=10/group: a) IgG control, b) anti-PD-1 (10 mg/kg, i.p., Q3Dx4), c) TGF-β inhibitor (Galunisertib, 75 mg/kg, p.o., QD), d) Combination.
- Tumor Measurement: Caliper measurements every 2 days. Volume = (length x width^2)/2.
- Endpoint Analysis: Day 28 harvest. Tumors dissociated for flow cytometry (anti-CD8, anti-FoxP3, anti-CD25). TILs quantified as % of live cells.
- Statistical Analysis: Two-way ANOVA for tumor growth; Log-rank test for survival.
Protocol 2: Surface-Modified CAR-T Functional Assay (Zhao & Li, 2024)
- CAR-T Engineering: Lentiviral transduction of primary human T-cells with mesothelin-targeting CAR construct incorporating a surface-tethered anti-PD-L1 scFv.
- Co-culture Assay: CAR-T cells co-cultured with PD-L1+ tumor spheroids (MIA PaCa-2) at 5:1 E:T ratio.
- Cytotoxicity: Measured via real-time cell analysis (RTCA) over 96h. Specific lysis = (1 - (impedance target+effector / impedance target alone)) x 100.
- Exhaustion Profiling: Day 5 CAR-T cells stained for PD-1, TIM-3, LAG-3. MFI measured via flow cytometry.
- In Vivo Validation: NSG mice with established tumors (>200 mm^3) treated with a single IV dose of 5x10^6 CAR-T cells. Bioluminescence imaging tracked persistence.
Signaling Pathways and Experimental Workflows
Diagram 1: Mechanism of TGF-β Inhibition Synergizing with ICB (76 chars)
Diagram 2: Workflow for Evaluating Combination Therapy Efficacy (78 chars)
Diagram 3: Surface-Modified CAR-T Dual-Action Mechanism (76 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Combination Therapy Research
| Reagent / Solution | Vendor Example (Catalog #) | Primary Function in Experiments |
|---|---|---|
| Recombinant Human TGF-β1 | PeproTech (100-21) | To stimulate TGF-β pathway in vitro for inhibitor validation assays. |
| Anti-Mouse PD-1 (Clone RMP1-14) | Bio X Cell (BE0146) | In vivo checkpoint blockade in syngeneic mouse models. |
| Galunisertib (LY2157299) | Selleckchem (S2230) | Small-molecule TGF-β receptor I kinase inhibitor for in vivo studies. |
| CellTrace Violet Proliferation Kit | Thermo Fisher (C34557) | To track T-cell or tumor cell division in co-culture experiments. |
| Mouse TIL Flow Panel (CD45, CD3, CD8, FoxP3, CD25) | BioLegend (Multiple) | Comprehensive immunophenotyping of the tumor microenvironment. |
| Human/Mouse TGF-β1 ELISA Kit | R&D Systems (DB100B/DB100C) | Quantify active TGF-β levels in serum or tumor lysates. |
| Lentiviral CAR Construct (MSLN scFv-41BB-CD3ζ) | VectorBuilder (Custom) | Generation of surface-modified CAR-T cells for functional assays. |
| GEM Hydrogel Tumor Organoid Kit | Cellendes (3D Life) | Establish 3D tumor spheroids for more physiologic cytotoxicity assays. |
Conclusion
Both direct TGF-β pathway inhibition and surface modification approaches offer powerful, yet distinct, routes to modulate this central biological pathway. Direct inhibitors provide broad, potent suppression but grapple with systemic toxicity and pathway pleiotropy. Surface modification strategies offer a more nuanced, localized, and potentially safer intervention, though they face hurdles in manufacturing and durability. The future lies in precision application: selecting the strategy based on disease context, stage, and tissue type. Emerging synergies, such as combining pathway inhibitors with surface-engineered cell therapies, represent a promising frontier. Ultimately, the choice is not between these modalities but in intelligently integrating them to overcome the TGF-β paradox and deliver transformative therapies for cancer and fibrosis.