Advanced CAD Design for Fully Interconnected Scaffold Channel Networks: Engineering the Future of Tissue Regeneration
This article provides a comprehensive guide for researchers, scientists, and drug development professionals on the cutting-edge CAD design of tissue engineering scaffolds with fully interconnected channel networks.
Advanced CAD Design for Fully Interconnected Scaffold Channel Networks: Engineering the Future of Tissue Regeneration
Abstract
This article provides a comprehensive guide for researchers, scientists, and drug development professionals on the cutting-edge CAD design of tissue engineering scaffolds with fully interconnected channel networks. We explore the fundamental biomechanical and biological principles underpinning scaffold architecture, detail the latest methodological workflows from concept to fabrication, address critical troubleshooting and optimization strategies for printability and nutrient flow, and finally, review validation techniques and comparative analyses against traditional designs. This systematic roadmap empowers the creation of next-generation scaffolds that maximize cell viability, vascularization potential, and functional tissue integration.
The Blueprint of Life: Foundational Principles for Interconnected Channel Design
Within the paradigm of computer-aided design (CAD) for tissue engineering scaffolds, the concept of a "fully interconnected" channel network is paramount. It is the cornerstone for ensuring uniform cell seeding, adequate nutrient/waste perfusion, and eventual vascularization in thick, clinically relevant constructs. This document establishes the quantitative and qualitative criteria that define the gold standard for full interconnectivity, providing application notes and protocols for researchers to validate their scaffold designs.
Defining Criteria for Full Interconnectivity
A channel network is considered fully interconnected when it satisfies the following geometric and topological parameters, measurable via CAD software and micro-CT analysis.
Table 1: Quantitative Criteria for a Fully Interconnected Channel Network
| Parameter | Definition & Measurement | Gold Standard Target Value | Functional Rationale |
|---|---|---|---|
| Connectivity Density (Conn.D) | Number of redundant connections per unit volume (mm⁻³). Measured via Euler-Poincaré characteristic. | > 10 mm⁻³ | Ensures multiple perfusion pathways, preventing failure from single blockages. |
| Global Porosity | Percentage of total scaffold volume occupied by void space (channels + microporosity). | 60 - 80% | Balances mechanical integrity with space for tissue ingrowth and flow. |
| Channel Interconnectivity (%) | Percentage of pore volume accessible from a single entrance. | 100% | No dead-end channels; all void space is perfusable. |
| Pore Throat Diameter | Minimum diameter of the connecting pathways between adjacent channel nodes. | ≥ 50 µm | Prevents cell bottlenecking and allows for capillary sprouting. |
| Tortuosity (τ) | Ratio of actual flow path length to straight-line distance. | < 2.0 | Low resistance to flow, promoting uniform medium/gradient distribution. |
| Surface Area to Volume Ratio | Total internal surface area per unit scaffold volume (mm²/mm³). | Scaffold-specific (e.g., 5-15 mm²/mm³) | Maximizes area for cell attachment while maintaining open channels. |
Protocol 1: CAD-Based Design Validation
Objective: To algorithmically verify full interconnectivity during the scaffold design phase. Workflow:
- Design Generation: Use CAD software (e.g., nTopology, SolidWorks, custom scripts) to generate a scaffold with an intended channel network (e.g., gyroid, orthogonal, or branched designs).
- Boolean Conversion: Convert the solid model into a 3D binary voxel dataset (volume element).
- Flood Fill Algorithm: Execute a 3D "flood fill" or "region growing" algorithm from a user-defined seed point within the void space.
- Analysis: Calculate the percentage of total void voxels reached by the algorithm.
- Pass: 100% of void voxels are accessed.
- Fail: <100% indicates the presence of isolated pores.
- Parameter Extraction: From the interconnected void volume, compute Conn.D, tortuosity, and pore throat diameters using image analysis libraries (e.g., scikit-image in Python).
Title: CAD Workflow for Interconnectivity Validation
Protocol 2: Experimental Validation via Micro-CT and Perfusion
Objective: To empirically validate the interconnectivity and permeability of a fabricated scaffold. Materials & Method: Part A: Structural Imaging (Micro-CT)
- Sample Preparation: Scan the fabricated scaffold (e.g., via 3D printing, decellularization) using high-resolution micro-CT (voxel size ≤ 1/3 of minimum pore throat diameter).
- Image Processing: Reconstruct 3D volume. Apply median filter, then global thresholding to segment scaffold material from void space.
- Morphological Analysis: Use BoneJ (ImageJ) or CTAN software to calculate parameters in Table 1 directly from the image stack. Confirm Conn.D > 10 mm⁻³ and 100% pore interconnectivity.
Part B: Functional Perfusion Test
- Setup: Mount the scaffold in a custom flow chamber or cartridge. Connect to a peristaltic pump and reservoir containing a tracer dye (e.g., Evans Blue) or fluorescent microbeads (Ø 10µm).
- Perfusion: Apply a constant low flow rate (e.g., 0.1 mL/min) to mimic interstitial flow. Capture the effluent at timed intervals.
- Analysis:
- Tracer Kinetics: Use a spectrophotometer/plate reader to measure tracer concentration in effluent vs. time. A smooth, sigmoidal uptake curve indicates uniform perfusion.
- Bead Distribution: Section the scaffold post-perfusion, image via fluorescence microscopy, and quantify bead distribution. Uniform distribution confirms functional interconnectivity.
Title: Experimental Validation Protocol Flow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Interconnectivity Research
| Item | Function & Relevance |
|---|---|
| CAD/CAE Software (nTopology, ANSYS) | Generates and simulates fluid flow through designed channel networks prior to fabrication. |
| High-Resolution 3D Printer (DLP, SLA) | Fabricates scaffolds with precise channel architectures (feature resolution < 50 µm). |
| Micro-CT Scanner (SkyScan, µCT) | Non-destructively images internal 3D microstructure for quantitative analysis. |
| Image Analysis Suite (BoneJ, CTAN) | Extracts critical 3D morphometric parameters (porosity, Conn.D, thickness) from image data. |
| Peristaltic Pump & Flow Chamber | Creates controlled perfusion conditions for functional testing of scaffold permeability. |
| Fluorescent Tracer Microbeads (Ø 2-20µm) | Act as cell-mimicking particles to visualize and quantify perfusion uniformity. |
| Biocompatible Hydrogels (GelMA, Alginate) | Used to infiltrate channels, assessing cell seeding efficiency and network occlusion. |
| Computational Fluid Dynamics (CFD) Software | Models shear stress, nutrient gradients, and pressure drops within the designed network. |
Within the thesis on CAD-designed scaffolds with fully interconnected channel networks, a foundational biological principle is paramount: three-dimensional interconnectivity is not a passive structural feature but a dynamic biological imperative. This document details the application notes and experimental protocols central to researching and validating this principle, focusing on cell viability, vascularization, and nutrient transport. The data underscores that pore and channel interconnectivity directly dictates metabolic survival, tissue ingrowth, and functional integration.
Application Notes: Quantitative Impact of Interconnectivity
Cell Viability and Metabolic Activity
Interconnected porosity prevents the formation of necrotic cores by facilitating waste removal and gas exchange. Isolated pores, regardless of size, lead to central hypoxia and apoptosis.
Table 1: Impact of Channel Interconnectivity on Cell Viability in 3D Constructs
| Interconnectivity Metric (Avg. Connections/Pore) | Max Viable Depth (µm) | Relative Glucose Consumption (Day 7) | Apoptotic Core % (Day 14) |
|---|---|---|---|
| <2 (Poorly Connected) | 150-200 | 0.45 ± 0.12 | 38.5 ± 5.2 |
| 3-4 (Moderately Connected) | 350-500 | 0.78 ± 0.09 | 12.1 ± 3.8 |
| >5 (Fully Connected Network) | >1000 | 1.00 ± 0.05 | 3.4 ± 1.5 |
Vascularization and Angiogenic Sprouting
Interconnected channels serve as physical guides for endothelial cell migration and tube formation. The degree of interconnection correlates with the speed and maturity of nascent vasculature.
Table 2: Vascularization Parameters in Scaffolds with Varied Interconnectivity
| Scaffold Type | Average Vessel Ingrowth Depth (mm, Week 2) | Branching Points/mm² | Perfused Vessel Fraction (%) |
|---|---|---|---|
| Non-Interconnected Porosity | 0.4 ± 0.1 | 15 ± 6 | <10 |
| Partially Interconnected Channels | 1.2 ± 0.3 | 42 ± 11 | 35-50 |
| CAD-Designed Full Network | 2.8 ± 0.5 | 85 ± 18 | 75-90 |
Nutrient and Biomolecule Transport
Convective flow and effective diffusion coefficients are exponentially enhanced by high interconnectivity, moving beyond simple diffusion-limited transport.
Table 3: Transport Efficiency Metrics
| Transport Mode | Effective Diffusion (Deff/D0) in Dense Cell Constructs | Convective Permeability (m²) |
|---|---|---|
| Diffusion-Only (No Channels) | 0.05 - 0.15 | N/A |
| Simple Parallel Channels | 0.3 - 0.4 | 1.2 x 10⁻¹² |
| Fully Interconnected Network | 0.6 - 0.8 | 5.8 x 10⁻¹² |
Detailed Experimental Protocols
Protocol 2.1: Quantifying Interconnectivity via Perfusion Analysis
Objective: To measure the functional interconnectivity of a 3D scaffold by determining its permeability to fluid flow. Materials: Scaffold sample, syringe pump, pressure transducer, PBS, tubing. Procedure:
- Mount the hydrated scaffold of known dimensions (e.g., 5mm dia x 2mm height) in a custom flow chamber.
- Connect the chamber inlet to a syringe pump and the outlet to a collection reservoir. Place a pressure transducer proximal to the inlet.
- Perfuse with PBS at incremental flow rates (Q: 0.1, 0.5, 1.0 mL/min).
- Record the steady-state pressure differential (ΔP) across the scaffold for each flow rate.
- Calculate permeability (κ) using Darcy’s Law: κ = (Q * μ * L) / (A * ΔP), where μ is fluid viscosity, L is scaffold thickness, and A is cross-sectional area.
- High permeability (κ > 1 x 10⁻¹² m²) indicates superior interconnectivity. Correlate κ values with cell viability data from identical scaffolds.
Protocol 2.2: Imaging and Assessing Viable Cell Depth
Objective: To spatially map live/dead cells within a seeded scaffold to determine the maximum depth of viability. Materials: Cell-seeded scaffold, Live/Dead Viability/Cytotoxicity Kit (calcein-AM/ethidium homodimer-1), confocal microscope, vibratome or cryosectioning setup. Procedure:
- Culture mesenchymal stem cells (MSCs) or relevant cell type on test scaffolds for 7-14 days.
- Rinse scaffolds in PBS and incubate in Live/Dead stain (2 µM calcein-AM, 4 µM EthD-1) for 45 minutes at 37°C.
- Image using a confocal microscope with z-stacking. Take orthogonal section views.
- For thick scaffolds (>1mm), optionally fix, embed, and section (e.g., 200 µm slices) using a vibratome before staining and imaging.
- Use image analysis software (e.g., FIJI/ImageJ) to calculate the depth from the nearest surface/channel at which the live cell fraction drops below 80%. This is the Max Viable Depth.
Protocol 2.3: In Vitro Angiogenesis Assay in Channeled Scaffolds
Objective: To quantify endothelial network formation within designed channel networks. Materials: HUVECs, fibrin or Matrigel, scaffolds, endothelial growth medium (EGM-2), angiogenic factors (VEGF, bFGF), confocal microscope. Procedure:
- Seed human umbilical vein endothelial cells (HUVECs) at 1 x 10⁶ cells/mL into the channel network of a fibronectin-coated scaffold using low-vacuum-assisted seeding.
- Fill the interstitial scaffold space with a fibrin gel (3 mg/mL) containing fibroblasts to provide trophic support.
- Culture in EGM-2 medium supplemented with 50 ng/mL VEGF and 30 ng/mL bFGF.
- At days 3, 7, and 14, fix samples and stain for CD31 (PECAM-1) and actin.
- Acquire 3D confocal image stacks. Analyze total tube length, number of branches, and network loops per unit volume using angiogenesis plug-ins (e.g., AngioTool).
Visualization of Key Pathways and Workflows
Diagram Title: Interconnectivity Drives Viability and Vascularization Pathways
Diagram Title: Scaffold Interconnectivity Validation Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Materials for Interconnectivity Research
| Item Name & Vendor Example | Function in Research | Critical Application Note |
|---|---|---|
| Live/Dead Viability/Cytotoxicity Kit (Thermo Fisher, L3224) | Simultaneously stains live (green, calcein-AM) and dead (red, EthD-1) cells. | Essential for Protocol 2.2. Use fresh stains and include a no-scaffold cell control for fluorescence baselines. |
| Matrigel Basement Membrane Matrix (Corning, 356231) | Provides a pro-angiogenic 3D environment for endothelial cells. | Used in Protocol 2.3. Keep on ice at all times before gelation to prevent premature polymerization. |
| Recombinant Human VEGF 165 (PeproTech, 100-20) | Key mitogen and chemoattractant for endothelial cells. | Critical for in vitro angiogenesis assays. Aliquot to avoid freeze-thaw cycles; use at 50-100 ng/mL. |
| CellTracker Deep Red Dye (Thermo Fisher, C34565) | Long-term, non-transferable cytoplasmic cell label for tracking migration. | Useful for visualizing cell infiltration depth into scaffold channels over time. |
| Fibrinogen from Bovine Plasma (Sigma, F8630) | Forms a tunable fibrin hydrogel to fill scaffold pores and support co-culture. | For Protocol 2.3. Combine with thrombin solution to gel around the seeded scaffold. |
| Anti-CD31/PECAM-1 Antibody (e.g., Abcam, ab28364) | Immunostaining marker for endothelial cells and nascent vasculature. | Use for quantifying vascular network formation in fixed samples from angiogenesis assays. |
| Pressure Transducer (e.g., Honeywell, 26PC Series) | Precisely measures pressure drop across a scaffold during perfusion. | Key for calculating permeability in Protocol 2.1. Calibrate against a known standard before use. |
Within the context of Computer-Aided Design (CAD) for scaffolds with fully interconnected channel networks, the precise balancing of porosity, pore size, and effective stiffness is paramount. This triad governs not only the mechanical integrity of the implant but also its biological performance, including nutrient diffusion, cell migration, proliferation, differentiation, and ultimately, tissue regeneration. This application note provides detailed protocols and synthesized data for researchers aiming to design and validate scaffolds where biomechanical properties are tuned via controlled architectural parameters.
Key Parameter Interrelationships & Quantitative Data
Table 1: Interdependency of Scaffold Architectural and Mechanical Properties
| Parameter | Typical Target Range for Bone Tissue Engineering | Influence on Permeability/Diffusion | Influence on Compressive Modulus | Primary CAD Control Method |
|---|---|---|---|---|
| Total Porosity | 60-90% | Exponential increase with porosity | Exponential decrease with porosity | Unit cell replication density and strut thickness. |
| Avg. Pore Size | 200-600 μm (bone) | Increases with pore size^2 (Hagen-Poiseuille) | Decreases with increasing pore size | Unit cell dimensions (e.g., cube edge length). |
| Pore Interconnectivity | >95% (fully interconnected) | Critical for uniform flow; limits dead zones | Minor effect if porosity is constant. | Lattice topology (e.g., gyroid vs. strut-based). |
| Effective Stiffness | 0.1-2 GPa (trabecular bone) | Indirect (via porosity relationship) | Direct design target via material and geometry. | Material assignment and minimal surface area. |
Table 2: Published Data on 3D-Printed PCL Scaffold Variants (Representative)
| Study Reference | Porosity (%) | Pore Size (μm) | Architecture | Compressive Modulus (MPa) | Key Cell Response Observation |
|---|---|---|---|---|---|
| Zein et al., 2002 | 60-80 | 400-800 | Fused deposition, orthogonal | 40-80 | Increased porosity enhanced osteoblast in-growth. |
| Hollister, 2005 | 50-70 | 400-500 | Image-based, gyroid | 10-50 | Stiffness and permeability predictable from CAD. |
| Giannitelli et al., 2015 | 70 | 500 | Salt-leached vs. 3D printed | 20 vs. 65 | Printed scaffolds showed superior mechanical stability. |
| Current CAD Benchmark | 75 ± 5 | 450 ± 50 | Triply Periodic Minimal Surface (TPMS) | 55 ± 15 | Optimal for MSC differentiation under perfusion. |
Experimental Protocols
Protocol 1: CAD-Based Design & Simulation of Interconnected Scaffolds
Objective: To generate a scaffold with defined porosity, pore size, and predicted stiffness using TPMS structures.
- Software: Utilize CAD (e.g., nTopology, Rhino3D with Grasshopper, or custom Python) to generate a Gyroid or Schwartz D unit cell.
- Parameterization: Define unit cell size (controls pore size) and volume fraction (controls porosity). E.g., for a 500μm pore size, use a ~1mm unit cell. Set volume fraction to 0.25 for ~75% porosity.
- Tessellation: Array the unit cell 5x5x5 times to create a bulk scaffold model.
- Boolean Operations: Intersect the tessellated lattice with a bounding geometry (e.g., a 10mm cylinder) to create the final implant shape.
- Export: Save the final design as an STL file for manufacturing and as a STEP file for simulation.
- Finite Element Analysis (FEA):
- Import STEP file into FEA software (e.g., ANSYS, Abaqus).
- Assign linear elastic material properties (e.g., PCL: E = 400 MPa, ν=0.3).
- Apply a compressive displacement (e.g., 2% strain) to one face while fixing the opposite face.
- Solve for stress and strain to calculate the effective elastic modulus (E_eff = stress / strain).
Protocol 2: Experimental Validation of Scaffold Permeability
Objective: To measure the Darcy permeability of a fabricated scaffold, validating interconnectivity.
- Setup: Use a custom or commercial permeability rig. Mount the sterilized scaffold in a water-tight chamber.
- Perfusion: Use a peristaltic pump to drive phosphate-buffered saline (PBS) or culture media through the scaffold at a constant flow rate (Q), ranging from 0.1 to 5 mL/min.
- Pressure Measurement: Record the pressure drop (ΔP) across the scaffold using in-line pressure sensors.
- Calculation: Apply Darcy's Law: K = (Q * μ * L) / (A * ΔP), where K is permeability (m²), μ is fluid viscosity (Pa·s), L is scaffold thickness (m), and A is cross-sectional area (m²).
- Analysis: Compare experimental K with CFD simulations of the original CAD model to confirm fabrication fidelity.
Protocol 3: Evaluating Cell Response to Biomechanical Cues in 3D Culture
Objective: To assess mesenchymal stem cell (MSC) differentiation in response to scaffold stiffness and pore architecture.
- Scaffold Preparation: Sterilize PCL scaffolds (varying stiffness/porosity) in 70% ethanol, rinse with PBS, and pre-wet in basal medium.
- Cell Seeding: Seed human MSCs at a density of 50,000 cells/scaffold using dynamic seeding (spinner flask) for 4 hours to ensure uniform penetration.
- Culture Conditions: Maintain in osteogenic medium (DMEM, 10% FBS, 10mM β-glycerophosphate, 50μM ascorbic acid, 100nM dexamethasone) under perfusion (0.5 mL/min) in a bioreactor for 21 days.
- Endpoint Analysis:
- Gene Expression (qPCR): Lyse cells, extract RNA, and analyze markers: Runx2 (early osteogenesis), OPN (mid), OCN (late).
- Histology: Fix scaffolds, section, and stain with Alizarin Red S (mineralization) and Hematoxylin & Eosin (cell distribution).
- Protein Synthesis (ELISA): Measure OPN and OCN secretion in conditioned media.
Diagrams
Design Parameter Impact on Osteogenesis
Scaffold Design-Validation Workflow
The Scientist's Toolkit: Research Reagent & Material Solutions
Table 3: Essential Materials for Scaffold Biomechanics Research
| Item | Function & Application | Example/Supplier |
|---|---|---|
| Polycaprolactone (PCL) | A biodegradable, FDA-approved polymer with tunable stiffness; ideal for fused filament fabrication (FFF) of scaffolds. | Sigma-Aldrich, 440744 |
| Triply Periodic Minimal Surface (TPMS) Design Software | Enables generation of mathematically defined, fully interconnected pore architectures with superior mechanical efficiency. | nTopology, Rhino3D (Grasshopper) |
| Perfusion Bioreactor System | Provides dynamic culture conditions to enhance nutrient/waste exchange and apply fluid shear stress in 3D scaffolds. | PBS Biotech, SQ-2 Series |
| Osteogenic Differentiation Kit | A defined, consistent supplement mix to induce and study osteoblast differentiation from progenitor cells. | Thermo Fisher, A1007201 |
| Micro-Computed Tomography (μCT) Scanner | For non-destructive 3D quantification of fabricated scaffold porosity, pore size, and interconnectivity. | Bruker, Skyscan 1272 |
| AlamarBlue Cell Viability Reagent | A resazurin-based assay for quantifying metabolic activity of cells within 3D scaffolds over time. | Thermo Fisher, DAL1100 |
| Human Mesenchymal Stem Cells (hMSCs) | Primary cells used to evaluate scaffold bioactivity and differentiation potential in regenerative medicine studies. | Lonza, PT-2501 |
Application Notes
Within the framework of CAD-driven design for tissue engineering scaffolds featuring fully interconnected channel networks, the triad of biocompatibility, degradation, and printability forms a critical design constraint loop. The channel network's primary function—to facilitate nutrient diffusion, waste removal, and potentially vascularization—is directly governed by these material properties.
Biocompatibility is non-negotiable and extends beyond baseline cytotoxicity. Materials must support specific cellular functions (e.g., adhesion, proliferation, differentiation) within the 3D channel-laden architecture. The high surface area of interconnected channels amplifies the material-cell interaction, making surface chemistry and degradation byproducts paramount. A biocompatible material that degrades into acidic monomers can locally alter pH in confined channels, adversely affecting encapsulated cells.
Degradation Rate must be engineered in lockstep with the CAD-designed geometry (e.g., strut thickness, channel diameter) and the intended tissue regeneration timeline. Bulk versus surface erosion modes dictate how channel patency and structural integrity are maintained. A mismatch, where the scaffold collapses before new tissue matrix is deposited, can occlude channels and lead to core necrosis. Synchronizing degradation with tissue ingrowth through the network is essential for mechanical and biological functionality.
Printability encompasses the rheological and physicochemical properties enabling the precise fabrication of complex, self-supporting channel networks (e.g., via extrusion-based or lithography-based bioprinting). Printability defines the fidelity of the CAD model to the physical construct, directly impacting channel interconnectivity, resolution, and surface topology. A highly biocompatible material with an ideal degradation profile is irrelevant if it cannot be printed into a robust, high-fidelity network.
These three factors are deeply interdependent. Adjusting material composition (e.g., polymer molecular weight, crosslink density) to tune degradation will alter melt viscosity or photocuring kinetics, affecting printability. Similarly, additives (e.g., bioceramics, plasticizers) included to enhance printability or biocompatibility can significantly modify the degradation profile.
Table 1: Common Biomaterials for 3D-Printed Scaffolds: A Triad Property Comparison
| Material Class & Example | Typical Biocompatibility Profile | Degradation Rate (Approx. Time for Mass Loss) | Key Printability Considerations |
|---|---|---|---|
| Synthetic Polymer (PCL) | Good; supports cell adhesion but relatively inert. | Slow; 2-4 years in vivo. Hydrolytic erosion. | Excellent for melt extrusion; low melting point (≈60°C), good viscoelasticity. |
| Synthetic Polymer (PLGA) | Good; widely used in FDA-approved devices. | Tunable (weeks to years); based on LA:GA ratio. Hydrolytic. | Suitable for extrusion (heating) and inkjet; viscosity control is critical. |
| Natural Polymer (Alginate) | Good; low immunogenicity but lacks cell-adhesive motifs. | Weeks to months; ion-dependent, can be rapid. | Excellent for extrusion-based bioprinting; ionotropic gelation enables crosslinking. |
| Natural Polymer (Gelatin Methacryloyl - GelMA) | Excellent; contains RGD sequences for cell adhesion. | Weeks to months; enzyme- and hydrolysis-dependent. | Premier for vat photopolymerization; photocrosslinkable, tunable modulus via concentration/UV. |
| Ceramic (β-Tricalcium Phosphate - β-TCP) | Excellent osteoconductivity; bioactive. | Slow; months to years; osteoclast-mediated resorption. | Printable via binder jetting or extrusion with polymers; often used in composites. |
| Composite (PCL/β-TCP, 70/30) | Enhanced osteoconductivity vs. PCL alone. | Slower than pure PCL; β-TCP buffers acidic PCL byproducts. | Enhanced stiffness vs. PCL; printability similar to PCL with optimized nozzle design. |
Experimental Protocols
Protocol 3.1: In Vitro Direct Contact Cytotoxicity Assay per ISO 10993-5 for Printed Scaffold Discs Purpose: To evaluate the baseline biocompatibility of a novel printable material formulation using scaffold discs with internal channel networks.
- Sample Preparation: Using CAD software, design a cylindrical scaffold (e.g., 5mm diameter x 2mm height) with a defined orthogonal channel network (e.g., 300µm channels). Print at least 24 identical discs using the candidate material under optimized parameters. Sterilize via ethylene oxide or ethanol immersion/UV.
- Cell Seeding: Culture L929 fibroblast cells or relevant primary cells in standard media. Seed cells into 24-well plates at 1 x 10^4 cells/well in 1 mL media and incubate for 24 hrs to allow attachment.
- Scaffold Application: Carefully place one sterile scaffold disc directly onto the cell monolayer in test wells. Use wells with cells alone as negative controls and wells with a polyurethane film containing 0.1% zinc diethyldithiocarbamate as a positive control.
- Incubation & Assessment: Incubate for 24-48 hours. Assess cell morphology microscopically. Perform a quantitative MTT assay: add MTT reagent (0.5 mg/mL), incubate 2-4 hrs, solubilize formazan crystals with DMSO, and measure absorbance at 570 nm.
- Analysis: Calculate cell viability relative to the negative control. A reduction in viability by >30% is considered a cytotoxic effect per ISO standards.
Protocol 3.2: Hydrolytic Degradation Profiling of Printed Scaffold Networks Purpose: To characterize mass loss, mechanical decay, and pH change of a degrading scaffold with interconnected channels.
- Scaffold Fabrication & Baseline: Print scaffold specimens (e.g., 10x10x3mm cubes with gyroid channel networks). Record dry mass (W0). Perform baseline compression testing (n=5) and SEM imaging of channel structure.
- Immersion Study: Immerse individual scaffolds (n=5 per time point) in 15 mL of phosphate-buffered saline (PBS, pH 7.4) at 37°C in sealed tubes. Include a blank PBS tube for pH monitoring.
- Time-Point Analysis: At predetermined intervals (e.g., 1, 2, 4, 8, 12 weeks): a. Remove scaffolds, rinse gently, dry under vacuum to constant weight (Wd). b. Measure pH of the incubation medium. c. Subject scaffolds to compression testing to determine retained modulus. d. For selected time points, image via SEM to observe channel morphology and surface erosion.
- Data Modeling: Plot % mass remaining [(Wd/W0)*100], compressive modulus, and pH vs. time. Fit mass loss data to kinetic models (e.g., zero-order, first-order) to predict degradation behavior.
Protocol 3.3: Printability & Fidelity Assessment for Interconnected Channel Designs Purpose: To quantitatively evaluate the capability of a bioink to reproduce a CAD-modeled channel network.
- Design & Printing: Create a benchmark CAD model featuring a 10x10x2mm lattice with two perpendicular, interconnected channel diameters (e.g., 400µm and 250µm). Print the model using the candidate bioink and optimized printer parameters.
- Fidelity Metrics: a. Macro-Fidelity: Use digital calipers to measure external dimensions (n=10). Calculate dimensional error vs. CAD. b. Channel Fidelity: Perform micro-CT scanning. Reconstruct the 3D model. Calculate: (i) Percent porosity vs. designed porosity; (ii) Channel diameter accuracy; (iii) Interconnectivity (% of designed channels that are fully patent).
- Rheological Assessment: Conduct oscillatory amplitude and frequency sweeps on the bioink to determine storage (G') and loss (G'') moduli, yield stress, and shear-thinning behavior—key predictors of extrudability and shape retention.
Diagrams
Diagram 1: Interdependent Design Loop for Scaffold Materials
Diagram 2: Workflow for Integrated Material Screening
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Scaffold Material Characterization
| Item | Function & Relevance |
|---|---|
| Gelatin Methacryloyl (GelMA) | A versatile, photocrosslinkable bioink derived from gelatin. Provides excellent biocompatibility (RGD motifs) and tunable mechanical/degradation properties, ideal for printing cell-laden channel networks. |
| Polycaprolactone (PCL), Medical Grade | A synthetic, biodegradable polyester with excellent thermal printability. Serves as a gold-standard material for studying the printing of complex, self-supporting channel architectures. |
| Photoinitiator (Lithium phenyl-2,4,6-trimethylbenzoylphosphinate - LAP) | A cytocompatible photoinitiator for UV/VL crosslinking of polymers like GelMA. Enables rapid gelation during printing to maintain channel shape fidelity. |
| Alginate (High G-Content) | A natural polysaccharide for ionic crosslinking (with Ca²⁺). Allows for gentle cell encapsulation and is a model material for studying degradation kinetics in channeled scaffolds. |
| β-Tricalcium Phosphate (β-TCP) Powder, <100nm | A bioactive ceramic used as an additive in composite bioinks to enhance osteoconductivity, modify degradation, and improve the mechanical strength of printed bone scaffolds. |
| Micro-Computed Tomography (Micro-CT) Scanner | Critical non-destructive equipment for 3D visualization and quantitative analysis of printed scaffold internal architecture, including channel interconnectivity, porosity, and wall thickness. |
| Rheometer with Peltier Plate | Essential for characterizing bioink viscoelasticity (storage/loss modulus, yield stress, shear-thinning) to predict and optimize printability for extrusion-based techniques. |
| MTT Assay Kit (ISO 10993-5) | Standardized colorimetric kit for quantifying in vitro cytotoxicity of material extracts or direct contact, providing a key biocompatibility metric. |
| Phosphate-Buffered Saline (PBS), pH 7.4 | Standard immersion medium for in vitro degradation studies, simulating physiological ionic strength and pH to monitor hydrolytic breakdown. |
Application Notes: Biomimetic Network Topologies in Scaffold Design
Thesis Context: This document supports research on Computer-Aided Design (CAD) for tissue engineering scaffolds with fully interconnected, biomimetic channel networks. The objective is to translate topologies observed in natural systems into CAD models that optimize nutrient diffusion, cell migration, and mechanical performance for drug screening and regenerative medicine applications.
Key Biomimetic Inspirations and Their Functional Analogies
Natural structures provide blueprints for optimal transport and structural efficiency. The following table summarizes quantitative characteristics of key bio-inspired topologies relevant to scaffold design.
Table 1: Quantitative Characteristics of Biomimetic Network Topologies for Scaffold Design
| Topology Type | Natural Inspiration | Key Geometric Parameters | Typical Porosity Range | Surface Area to Volume Ratio | Relative Permeability/Diffusivity | Mechanical Stiffness (Relative) |
|---|---|---|---|---|---|---|
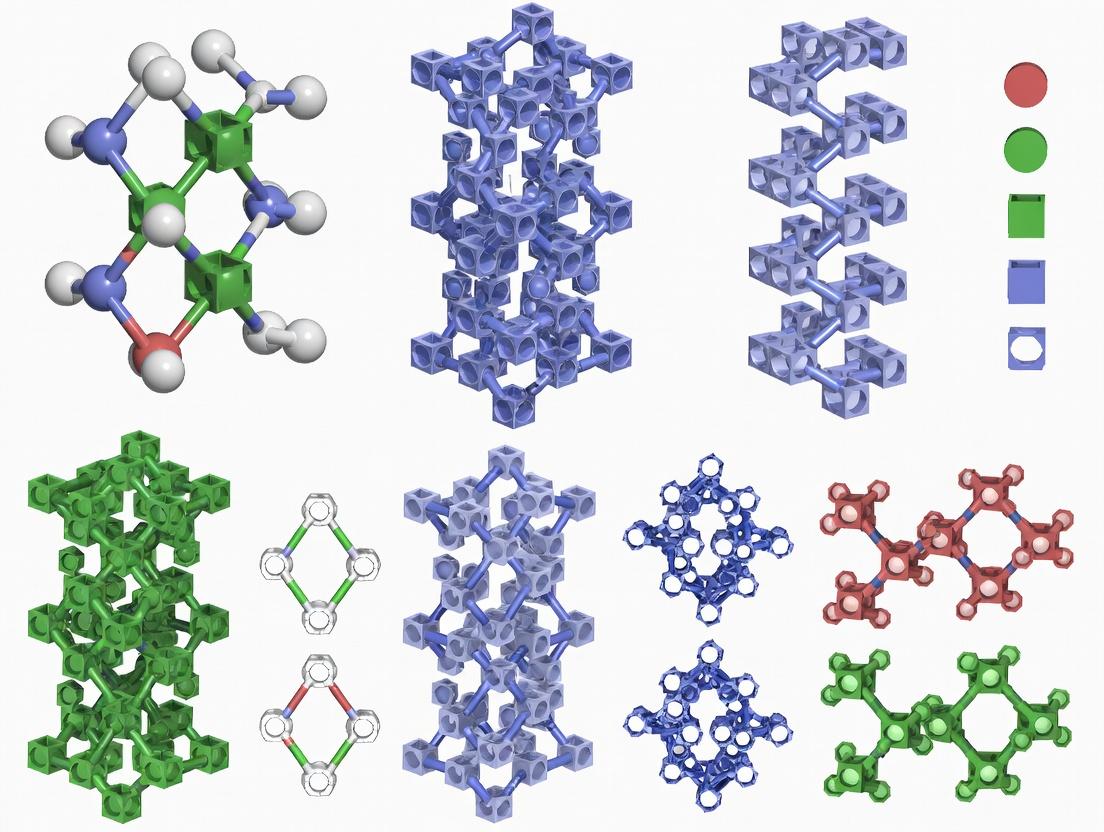
| Triply Periodic Minimal Surfaces (TPMS) - Gyroid | Butterfly wing scales, sea urchin skeletons | Unit cell size (μm), Wall thickness (μm), Porosity (%) | 50% - 80% | Very High (~2-5x strut-based lattices) | Excellent (intertwined, interconnected channels) | High (isotropic, smooth curvature) |
| Voronoi Structures | Trabecular bone, cork, foam structures | Seed point density (#/mm³), Cell size variance (CV%), Mean edge length (μm) | 60% - 90% | Moderate to High | Good (dependent on window connectivity) | Variable (can mimic bone stiffness gradient) |
| Leaf Venation Patterns | Plant leaves (e.g., dicotyledons) | Channel diameter hierarchy (primary: >100μm, tertiary: <20μm), Branching angle (deg) | N/A (planar) | High (planar) | Excellent for directed flow | Low (planar, often 2.5D) |
| Lattice Networks (Strut-based) | Honeycomb, coral | Strut diameter (μm), Node connectivity, Pore size (μm) | 70% - 95% | Moderate | Good (can be anisotropic) | Very High (for given porosity) |
Performance Metrics for Interconnectedness
A fully interconnected pore network is critical for cell viability and vascularization. Key metrics derived from computational analysis include:
Table 2: Metrics for Assessing Channel Network Interconnectedness
| Metric | Definition | Measurement Method (Typical) | Target Value for Scaffolds |
|---|---|---|---|
| Connectivity Density (CD) | Number of redundant connections per unit volume. | Micro-CT analysis, Euler number calculation. | >10 mm⁻³ |
| Percent Interconnectivity | Volume fraction of pores connected to the main network. | Image analysis via pore labeling. | >99.5% |
| Tortuosity (τ) | Ratio of actual flow path length to straight-line distance. | Computational fluid dynamics (CFD) simulation. | 1.2 - 2.5 (lower enhances diffusion) |
| Pore Access Size | Diameter of the largest sphere that can traverse the network. | Morphological opening algorithm on 3D model. | >30μm (for cell migration) |
Experimental Protocols
Protocol: Computational Generation and Analysis of Biomimetic Topologies
Aim: To generate, mesh, and simulate fluid flow through bio-inspired CAD models for scaffold evaluation.
Materials & Software:
- CAD/Modeling Software (e.g., nTopology, Rhino3D with Grasshopper, MATLAB)
- Finite Element Analysis (FEA)/CFD Software (e.g., COMSOL Multiphysics, ANSYS Fluent)
- High-Performance Computing (HPC) workstation.
Methodology:
- Parametric Model Generation:
- Gyroid: Define using the implicit function:
sin(x)*cos(y) + sin(y)*cos(z) + sin(z)*cos(x) = t. Vary the thresholdtto control porosity. - Voronoi: Generate random seed points within a volume. Compute 3D Voronoi tessellation. Convert edges to cylindrical struts or use cell walls as surfaces.
- Conformal Lattices: Create unit cell (e.g., diamond, octet-truss). Array unit cells in 3D space. Apply conformal smoothing to node junctions to reduce stress concentrations.
- Gyroid: Define using the implicit function:
- Mesh Export: Export the solid geometry as a watertight STL or STEP file.
- CFD Simulation for Permeability:
- Import geometry into CFD software.
- Define fluid properties (e.g., culture medium viscosity).
- Apply a pressure gradient (ΔP) across the scaffold model.
- Solve the steady-state Navier-Stokes equations for incompressible flow.
- Calculate permeability (k) using Darcy's Law:
k = (Q * μ * L) / (A * ΔP), where Q is volumetric flow rate, μ is viscosity, L is scaffold length, A is cross-sectional area.
- Mechanical Simulation:
- Assign isotropic material properties (e.g., PLA, PCL, Ti-6Al-4V) to the geometry.
- Apply a fixed boundary condition on one face and a uniaxial displacement/load on the opposite face.
- Solve for stress and strain fields to determine effective elastic modulus.
Protocol: Physical Fabrication & Interconnectivity Validation via Micro-CT
Aim: To fabricate biomimetic scaffolds via additive manufacturing and quantitatively validate channel network interconnectivity.
Materials:
- Biocompatible polymer resin (e.g., PEGDA) or metal powder (Ti-6Al-4V).
- Additive manufacturing system (e.g., Digital Light Processing (DLP) printer, Selective Laser Melting (SLM)).
- Micro-Computed Tomography (Micro-CT) system (e.g., SkyScan 1272).
- Image analysis software (e.g., CTAn, ImageJ/Fiji).
Methodology:
- Fabrication:
- Convert final CAD model to machine-specific build file (e.g., .slc for DLP, .slm for SLM).
- For polymers: Print using DLP with 405nm light, layer thickness 25-50μm. Post-process: rinse in IPA, post-cure under UV light.
- For metals: Print using SLM under argon atmosphere with optimized laser power and scan speed.
- Micro-CT Scanning:
- Mount scaffold securely on stage.
- Set scanning parameters: Voltage 40-80 kV, Current 100-200 μA, Pixel Size 3-10μm (to resolve pore features), Rotation step 0.4-0.7°, 180° or 360° rotation.
- Acquire projection images.
- Image Reconstruction & Analysis:
- Reconstruct 2D projections into a 3D volume (stack of cross-sections) using filtered back-projection.
- Binarization: Apply a global threshold (e.g., Otsu's method) to segment scaffold material from pores/channels.
- Quantification:
- Calculate total porosity (
Po(tot)). - Perform a "3D object labeling" analysis on the pore phase. The largest connected pore cluster is defined as the accessible network. Calculate Percent Interconnectivity as:
[Volume of Largest Pore Cluster / Total Pore Volume] * 100. - Perform a "sphere fitting" analysis to calculate pore size distribution and access size.
- Calculate total porosity (
Visualization: Pathway for Biomimetic Scaffold Design & Evaluation
Design & Evaluation Workflow for Biomimetic Scaffolds
The Scientist's Toolkit: Key Research Reagent Solutions & Materials
Table 3: Essential Materials for Biomimetic Scaffold Research
| Item Name | Category | Function / Relevance | Example Supplier/Product |
|---|---|---|---|
| Poly(ethylene glycol) diacrylate (PEGDA) | Photopolymer Resin | A biocompatible, photocurable resin for DLP printing of hydrogel scaffolds. Allows tuning of mechanical properties via molecular weight. | Sigma-Aldrich, 701963 |
| Ti-6Al-4V ELI Powder | Metal Feedstock | Grade 23 titanium alloy powder for SLM printing of high-strength, osteoconductive bone scaffolds. | AP&C (GE Additive), 15-45μm spherical powder |
| Iridium Platinum (Ir/Pt) Sputter Coater | Sample Preparation | Provides a thin, conductive coating on non-conductive polymer scaffolds for high-quality SEM imaging without charging artifacts. | Quorum Technologies, Q150T S |
| AlamarBlue Cell Viability Reagent | Biological Assay | Resazurin-based dye used to assess metabolic activity of cells seeded on 3D scaffolds, indicating cytocompatibility. | Thermo Fisher Scientific, DAL1100 |
| Matrigel Basement Membrane Matrix | Hydrogel/Cell Carrier | Used to coat scaffold interiors or embed cells to enhance cell attachment, proliferation, and formation of 3D structures within channels. | Corning, 356231 |
| Micro-CT Calibration Phantom | Imaging Standard | A phantom with known density standards (e.g., hydroxyapatite) is scanned alongside samples to calibrate mineral density quantification in bone tissue engineering studies. | Bruker, Morphology Phantom |
| ImageJ/Fiji with BoneJ Plugin | Image Analysis Software | Open-source platform for analyzing 3D micro-CT data (porosity, thickness, connectivity) and quantifying scaffold morphology. | Open Source (NIH) |
From Pixels to Biomaterial: A Step-by-Step CAD Methodology for Complex Channel Networks
Application Notes and Protocols
This review is framed within the context of a doctoral thesis on CAD design for tissue engineering scaffolds with fully interconnected, perfusion-enhancing channel networks. The focus is on evaluating software capabilities for generating, optimizing, and validating complex, biomimetic architectures for in vitro and in vivo applications in drug development and regenerative medicine.
Core Software Comparison for Scaffold Design
The following table summarizes key quantitative and qualitative metrics for the primary software toolkits, based on current specifications, documentation, and research applications.
Table 1: Comparative Analysis of Software for Interconnected Channel Scaffold Design
| Software | Primary Strength | Lattice/TPMS Generation | Native TO | Interconnectivity Assurance | Bio-export Formats (STL, 3MF, STEP) | Learning Curve | Approx. Cost (Academic) |
|---|---|---|---|---|---|---|---|
| nTopology | Implicit modeling & field-driven design | Excellent (Custom & TPMS) | Advanced (Lattice, Density) | Built-in (Boolean & Field ops) | STL, 3MF, STEP | Moderate to High | $2,500 - $5,000/yr |
| SolidWorks | Parametric solid modeling | Basic (Add-ins) | Basic (Simulation Premium) | Manual assembly control | STL, 3MF, STEP | Moderate | ~$3,500/yr |
| Autodesk Netfabb | Additive Manufacturing prep & lattice | Good (TPMS via tools) | Good (Local Lat. TO) | Via analysis tools | STL, 3MF, AMF | Moderate | ~$1,600/yr (Suite) |
| Rhino/Grasshopper | Flexible NURBS & algorithmic modeling | Excellent (Plugins: Pufferfish, Intralattice) | Good (Plugins: Millipede, TopOpt) | Algorithmically defined | STL, 3MF, STEP | High (for GH) | ~$995 (Rhino) |
Experimental Protocols for Software Evaluation in Thesis Research
Protocol 1: Benchmarking Channel Network Interconnectivity
- Objective: Quantify the software's ability to generate and verify a fully interconnected pore/channel network suitable for cell seeding and perfusion.
- Materials:
- Workstation (High RAM >32GB, multi-core CPU)
- Target Software (as per Table 1)
- Validation software (e.g., VoxelPrint, simple Python script for flood-fill algorithm)
- Methodology:
- Model Generation: In each software, design a 10x10x10 mm scaffold block with a defined triply periodic minimal surface (TPMS) – Gyroid unit cell. Target pore size: 500 µm, strut thickness: 200 µm.
- Boolean Subtraction: Subtract the TPMS lattice from a solid block to create the porous scaffold. In nTop, use implicit Boolean. In SolidWorks, use solid subtract. In Netfabb, use lattice tool. In Grasshopper, bake and Boolean.
- Export: Export the model in STL format at a resolution of 0.05 mm.
- Interconnectivity Validation:
a. Convert STL to a binary voxel grid (e.g., using Python
numpy-stlandscikit-image). b. Apply a 3D flood-fill algorithm from a central seed voxel. c. Calculate the percentage of scaffold void volume filled. 100% indicates full interconnectivity. - Measurement: Record time-to-design, file size, and computational load for each software.
Protocol 2: Topology Optimization for Mechanical and Permeability Goals
- Objective: Generate a scaffold topology that meets both mechanical stiffness under a compressive load and maximizes fluid permeability (a surrogate for nutrient diffusion).
- Materials: Software with TO capabilities (nTopology, Netfabb, Grasshopper+Millipede), FEA solver (integrated or Abaqus).
- Methodology:
- Define Design Space: A cylindrical volume (Ø8mm x 10mm height).
- Load Cases & Constraints: Fix bottom surface. Apply 1MPa distributed compressive load on top surface.
- Objective & Constraints: Maximize permeability (modeled as minimizing pressure drop via Darcy flow simulation in nTop or via post-processing) while constraining global compliance (stiffness) to be ≤ 2x that of a solid PLA model. Target a 70% volume fraction.
- Optimization Execution: Run the multi-physics TO. In nTop, use the
Field-Driven Optimizationblock. In Grasshopper, use Millipede's multi-objective setup. - Post-Processing: Smooth the result, remesh for manufacturability, and run verification FEA/CFD.
Visualization: Software Selection and Validation Workflow
Title: Workflow for Scaffold Design Software Evaluation
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Materials and Digital Tools for Scaffold Design & Validation
| Item / Software Module | Function in Research | Example/Supplier |
|---|---|---|
| TPMS Algorithm Library | Generates mathematically defined, smooth, interconnected pore architectures. Essential for biomimetic design. | nTopology Implicit Body, Grasshopper Pufferfish plugin, MSLattice. |
| Voxel-Based Analysis Tool | Converts surface mesh (STL) to voxel grid for computational analysis of porosity, connectivity, and permeability. | Python (scikit-image, pyvista), VoxelPrint, ImageJ/Fiji. |
| Lattice Optimization Engine | Optimizes lattice cell size, thickness, or density distribution to meet mechanical targets while preserving channels. | nTopology Lattice Optimization, Netfabb Local Lat. TO, Altair Inspire. |
| Multi-Physics Solver | Simulates coupled physical phenomena (e.g., fluid-structure interaction) to predict scaffold performance under bioreactor conditions. | COMSOL, ANSYS Fluent/Mechanical, nTopology Field Analysis. |
| High-Resolution STL/3MF Exporter | Prepares final digital model for additive manufacturing (e.g., SLA, DLP, 2PP) with critical mesh integrity. | Native exports from all reviewed software; MeshLab for repair. |
| Biocompatible Resin | Material for physical prototype fabrication. Must be suitable for cell culture (e.g., Class VI, or biodegradable). | Formlabs BioMed, PEGDA-based resins, Polycaprolactone (PCL) filaments. |
Within the broader research on CAD design for scaffolds with fully interconnected channel networks, the primary objective is to engineer porous architectures that precisely control mass transport (e.g., nutrients, oxygen, metabolites) and cellular infiltration. This is critical for tissue engineering and in vitro drug testing models. Traditional design methods are limited in exploring the vast design space for optimal channel topologies. This document details application notes and protocols for a generative design workflow that employs algorithms to automatically create and refine channel paths, ensuring full interconnectivity and meeting specific biological and mechanical constraints.
Generative workflows for channel networks typically leverage space colonization, reaction-diffusion, or voronoi-based algorithms, followed by computational fluid dynamics (CFD) and finite element analysis (FEA) for evaluation.
Table 1: Comparative Analysis of Generative Algorithms for Channel Pathing
| Algorithm | Key Principle | Primary Output | Typical Porosity Range (%) | Computational Cost | Optimal Use Case |
|---|---|---|---|---|---|
| Space Colonization | Growth from seed points towards target points, avoiding occupied space. | Tree-like, branched networks. | 60-85 | Low-Moderate | Mimicking vascular or neuronal branching structures. |
| Voronoi Tessellation | Partitioning space based on distance to seed points. | Stochastic, polyhedral pore networks. | 70-90 | Low | Creating biomimetic, foam-like architectures. |
| Reaction-Diffusion (e.g., Murray's Law) | Modeling morphogen gradients to dictate branch diameter and bifurcation. | Physiologically optimized fluidic networks. | 50-75 | Moderate-High | Engineering vascular networks for optimal shear stress and flow. |
| Lattice Boltzmann Method (LBM) Optimization | Simulating fluid flow to iteratively erode/add material. | Pressure-drop optimized paths. | 65-80 | Very High | Maximizing perfusion efficiency in thick scaffolds. |
| Triply Periodic Minimal Surfaces (TPMS) | Mathematical implicit functions (e.g., Gyroid, Schwarz D). | Smooth, highly interconnected surfaces. | 40-70 | Moderate | Scaffolds with superior mechanical strength and mixed convection-diffusion transport. |
Table 2: Key Performance Metrics for Algorithmic Channel Networks
| Performance Metric | Target Range (Tissue Engineering Scaffold) | Analysis Method | Typical Benchmark Value for Generative Designs |
|---|---|---|---|
| Interconnectivity (%) | 100% (Fully Interconnected) | Micro-CT analysis, pore connectivity index. | >99.5% |
| Wall Shear Stress (Pa) | 0.1 - 3.0 Pa (for endothelial cells) | Computational Fluid Dynamics (CFD). | 0.5 - 2.5 Pa (optimized networks) |
| Permeability (m²) | 10⁻¹⁰ - 10⁻⁸ | CFD via Darcy's Law. | 5.0 x 10⁻¹⁰ |
| Diffusion Efficiency | Maximized | Simulation of molecular diffusion. | >30% improvement vs. random pores. |
Experimental Protocols
Protocol 3.1: Generative Workflow for a Perfusable Branched Network Objective: To algorithmically generate a Murray's Law-optimized branched channel network within a cubic scaffold for subsequent fabrication and perfusion culture. Materials: Workstation with Python/R/MATLAB, CAD software (e.g., Rhino 3D with Grasshopper), CFD software (e.g., ANSYS Fluent, COMSOL).
Algorithmic Generation (Python Script Example):
- Seed Definition: Define inlet and outlet boundary coordinates within a 10x10x10 mm design volume.
- Space Colonization Execution: Implement algorithm where growth nodes iteratively move towards randomly distributed target points within the volume, with a defined kill distance.
- Diameter Assignment: Apply Murray's Law (dparent³ = dchild1³ + d_child2³) at each bifurcation to calculate daughter branch diameters, assuming a constant bifurcation exponent (n=3).
- Mesh Generation: Convert the resultant node-and-branch data into a 3D cylindrical network with smooth Boolean unions. Export as an STL file.
CFD Validation:
- Mesh Import & Preparation: Import the STL into CFD pre-processor. Generate a volumetric mesh around the channel geometry.
- Boundary Conditions: Set inlet as velocity inlet (typical 1 mm/s for interstitial flow). Set outlets as pressure outlets (0 Pa).
- Solver Setup: Use a laminar flow model. Set fluid properties to mimic culture media (density: 1000 kg/m³, viscosity: 0.001 Pa·s).
- Simulation & Analysis: Solve for velocity and pressure fields. Extract wall shear stress (WSS) data across all channel surfaces. Verify WSS is within the target biological range (0.1-3.0 Pa).
Design Refinement Loop:
- Identify channels with WSS outside the target range.
- Adjust the algorithm's parameters (e.g., kill distance, branch attraction force, minimum diameter constraint) to thicken (for low WSS) or thin (for high WSS) specific branches.
- Re-run the generative algorithm and CFD simulation iteratively until WSS criteria are met.
Protocol 3.2: Experimental Validation of Channel Interconnectivity via Micro-CT Objective: To verify the physical interconnectivity of an additively manufactured scaffold generated via the above workflow. Materials: Fabricated scaffold (e.g., via stereolithography), micro-CT scanner (e.g., SkyScan 1272), image analysis software (e.g., CTAn, ImageJ).
- Sample Preparation: Mount the scaffold securely on the specimen stage. Ensure no movement during rotation.
- Scanning Parameters: Set voltage to 50 kV, current to 200 µA. Use a 0.5 mm aluminum filter. Set voxel size to 5 µm (sufficient to resolve channel walls). Perform a 180° rotation with a 0.4° rotation step.
- Image Reconstruction: Use the scanner's proprietary software (e.g., NRecon) to reconstruct projection images into cross-sectional slices. Apply consistent beam hardening and ring artifact correction.
- Interconnectivity Analysis (CTAn Software):
- Thresholding: Apply a global threshold to binarize images into solid material and pore space.
- 3D Analysis: Run the "Analysis" function to calculate total porosity and closed porosity.
- Pore Connectivity: Execute the "Pore Connectivity" plugin. This labels all interconnected pore voxels.
- Calculation: Interconnectivity (%) = [(Total Porosity - Closed Porosity) / Total Porosity] * 100. A result >99.5% confirms a fully interconnected network.
Visualizations
Diagram 1: Generative Design & Validation Workflow
Diagram 2: Space Colonization Algorithm Logic
The Scientist's Toolkit: Research Reagent Solutions & Essential Materials
Table 3: Essential Computational & Experimental Tools
| Item / Reagent | Function / Purpose | Example / Notes |
|---|---|---|
| Generative Scripting Environment | Core platform for implementing and customizing design algorithms. | Python with NumPy, SciPy libraries; Rhino 3D Grasshopper with plugins like Anemone. |
| Computational Fluid Dynamics (CFD) Software | Simulating fluid flow, shear stress, and diffusion within designed channels. | ANSYS Fluent, COMSOL Multiphysics, OpenFOAM (open-source). |
| High-Resolution 3D Printer | Physically fabricating the algorithmically generated scaffold designs. | Stereolithography (SLA) printers (e.g., Formlabs) for <100 µm features; Two-Photon Polymerization (2PP) for sub-micron resolution. |
| Micro-Computed Tomography (Micro-CT) System | Non-destructive 3D imaging to quantify porosity, interconnectivity, and channel fidelity. | Bruker SkyScan 1272; Critical for Protocol 3.2. |
| Image Analysis Suite | Processing 3D image data from micro-CT to extract quantitative metrics. | Bruker CTAn, ImageJ/Fiji with BoneJ plugin. |
| Biocompatible Photopolymer Resin | Material for fabricating scaffolds intended for biological validation. | Formlabs Biomedical Resin, Polyethylene glycol diacrylate (PEGDA)-based resins. |
| Perfusion Bioreactor System | Experimental validation of channel network functionality under dynamic culture. | Custom or commercial systems (e.g., IBIDI Pump System) to apply physiological flow rates. |
This application note details the parametric modeling protocols developed for a broader doctoral thesis on "CAD-Driven Design of Biphasic Scaffolds with Fully Interconnected Channel Networks for Osteochondral Tissue Engineering." The core objective is to establish a robust, adaptable Computer-Aided Design (CAD) framework that enables the precise and independent control of three critical scaffold architectural parameters: pore size, channel diameter, and wall thickness. This control is fundamental to optimizing mechanical properties, nutrient diffusion, cell seeding efficiency, and ultimately, tissue regeneration within the biphasic construct.
Key Quantitative Parameters & Design Targets
Table 1: Target Parameter Ranges for Osteochondral Scaffold Design
| Architectural Parameter | Target Range (µm) | Phase Association | Primary Biological Function |
|---|---|---|---|
| Pore Size | 200 - 500 | Cartilaginous Phase | Chondrocyte attachment & ECM production |
| Channel Diameter | 500 - 1000 | Osseous Phase | Vascularization & bone ingrowth |
| Wall Thickness | 100 - 300 | Both Phases | Mechanical integrity & degradation kinetics |
Core Parametric Modeling Protocol
Software & Environment Setup
- Primary CAD Software: Dassault Systèmes SolidWorks 2024 or Siemens NX 2306.
- Key Tools: Equation-driven curves, pattern features, configurations, and design tables.
- File Management: Maintain a master part file with all global variables and linked derived documents.
Step-by-Step Modeling Workflow
Step 1: Define Global Variables. Initiate the model by declaring global driving variables.
Step 2: Generate Unit Cell Lattice. Create a 2D sketch on the front plane using equation-driven curves to form a repeating unit (e.g., gyroid, Schwarz diamond, or custom truncated octahedron). Dimension all sketch entities by linking to the global variables.
Step 3: Extrude to 3D Solid & Pattern. Extrude the sketch to a depth defined by Unit_Cell_Size. Use a linear pattern feature in X, Y, and Z directions, spacing set to Unit_Cell_Size, to create a 5x5x5 lattice block as the base scaffold volume.
Step 4: Create Interconnected Channels.
- Define a new plane at the center of the lattice block.
- Sketch the channel network (orthogonal or staggered). Dimension channel circles/rectangles using the
Channel_Diavariable. - Use the "Combine" or "Subtract" command with the "Save Tool Bodies" option to subtract the channel sweep from the lattice, creating the final interconnected network. This step is performed separately for osseous and cartilaginous phase models.
Step 5: Implement Configurations for Design Exploration. Utilize the Configurations feature to create multiple design variants within a single file. A Design Table (Excel spreadsheet linked to the CAD file) is populated to manage variants systematically.
Table 2: Example Design Table for Configuration Management
| Configuration Name | Pore_Size (µm) | Channel_Dia (µm) | Wall_Thickness (µm) | Porosity (%) |
|---|---|---|---|---|
| Design_V1 | 200 | 500 | 100 | ~78% |
| Design_V2 | 350 | 750 | 150 | ~82% |
| Design_V3 | 500 | 1000 | 200 | ~85% |
Step 6: Export for Manufacturing & Simulation. Export each configuration as an STL file for additive manufacturing (e.g., stereolithography, selective laser sintering) or as a STEP file for finite element analysis (FEA) in software like ANSYS or COMSOL.
Experimental Validation Protocol for Printed Scaffolds
Micro-Computed Tomography (µCT) Characterization
- Objective: Quantify actual pore size, channel diameter, wall thickness, and interconnectivity.
- Protocol:
- Scan: Scan scaffold (n=5 per design) using a Skyscan 1272 system at 5 µm resolution, 60 kV, 166 µA.
- Reconstruction: Use NRecon software with standardized beam hardening and ring artifact correction.
- Analysis: Import reconstructed data into CTAn software. Apply a fixed global threshold to binarize images. Calculate porosity, pore size distribution (Sphere Fitting method), and structure thickness (3D Thickness plugin).
Uniaxial Compression Testing
- Objective: Correlate parametric design with mechanical performance.
- Protocol:
- Conditioning: Hydrate scaffolds in PBS for 24h at 37°C.
- Testing: Perform test on Instron 5944 with a 1 kN load cell at a strain rate of 0.5 mm/min until 50% strain.
- Calculation: Determine compressive modulus from the linear region of the stress-strain curve (typically 2-10% strain).
Visualization of the Parametric Design & Validation Workflow
Diagram Title: Parametric CAD to Physical Validation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Scaffold Fabrication & Analysis
| Item Name | Supplier (Example) | Function/Application |
|---|---|---|
| Polycaprolactone (PCL) | Sigma-Aldrich, 440744 | Synthetic polymer for fused deposition modeling (FDM); provides tunable mechanical strength and slow degradation. |
| Tricalcium Phosphate (TCP) Powder | Berkeley Advanced Biomaterials, <20µm | Bio-ceramic filler for composite printing; enhances osteoconductivity in the osseous phase. |
| GelMA (Gelatin Methacryloyl) | Advanced BioMatrix, GEL-100 | Photocrosslinkable bioink for stereolithography; forms the hydrog el-like cartilaginous phase. |
| Lithium Phenyl-2,4,6-trimethylbenzoylphosphinate (LAP) | Sigma-Aldrich, 900889 | Efficient photo-initiator for UV crosslinking of GelMA and similar polymers. |
| Phosphate Buffered Saline (PBS), 10X | Thermo Fisher Scientific, 70011044 | Standard buffer for scaffold hydration, washing, and as a cell culture medium supplement. |
| AlamarBlue Cell Viability Reagent | Thermo Fisher Scientific, DAL1025 | Resazurin-based solution for non-destructive, quantitative assessment of cell proliferation on scaffolds. |
| Fluorescein Diacetate (FDA) / Propidium Iodide (PI) | Sigma-Aldrich, F7378 / P4170 | Live/Dead viability staining kit for direct visualization of cell distribution and viability within 3D scaffolds. |
This Application Note details protocols for translating CAD models of scaffolds with fully interconnected channel networks into physical constructs via three dominant bioprinting modalities: Stereolithography (SLA), Digital Light Processing (DLP), and Extrusion-based bioprinting. Within the broader thesis on CAD for vascularized tissue engineering, manufacturability is the critical bridge between computational design (e.g., topology-optimized, gyroid, or branching channel networks) and biologically functional scaffolds. The primary challenges addressed herein are model preparation, material constraints, and print parameter optimization to ensure channel patency, shape fidelity, and biocompatibility.
Quantitative Comparison of Bioprinting Modalities for Channel Fabrication
Table 1: Key Quantitative Parameters for Channel Network Fabrication Across Bioprinting Technologies.
| Parameter | SLA | DLP | Extrusion-based |
|---|---|---|---|
| Typical XY Resolution | 50 - 150 µm | 20 - 50 µm | 100 - 500 µm |
| Typical Z-Layer Height | 25 - 100 µm | 10 - 50 µm | 50 - 300 µm |
| Minimum Viable Channel Diameter | 150 - 200 µm | 50 - 100 µm | 200 - 400 µm |
| Typical Viscosity Range | 0.5 - 5 Pa·s | 0.5 - 5 Pa·s | 30 - 10⁶ Pa·s |
| Critical Print Speed | 10 - 100 mm/s (laser scan) | 1 - 10 s/layer (layer cure) | 1 - 20 mm/s (nozzle speed) |
| Key Channel Occlusion Risk | Laser over-cure, resin swelling | Light scattering, over-penetration | Nozzle pressure collapse, filament fusion |
| Post-processing Requirement | Solvent rinse, UV post-cure | Solvent rinse, UV post-cure | Incubation for crosslinking |
Experimental Protocols for Model Preparation & Print Validation
Protocol 3.1: Universal Pre-Print CAD Preparation for Interconnected Channels
- Design Import: Import scaffold model (e.g., .STL, .OBJ) into mesh repair software (e.g., Autodesk Meshmixer, Netfabb).
- Mesh Analysis & Repair: Run automated repair to fix non-manifold edges, self-intersections, and inverted normals. Manually inspect and seal unintended openings in the external geometry only.
- Channel Patency Verification:
- Use software's "inspector" tool to visually confirm channel inlets/outlets.
- Employ a virtual "sphere test" (simulate a sphere of diameter equal to 80% of minimum designed channel width traversing the network).
- Support Structure Generation (for SLA/DLP):
- Set support contact tip diameter to 0.2-0.3 mm.
- Angle supports ≥ 45° from the build platform to minimize contact with critical channel surfaces.
- Export supported model in printer-specific slice file format (e.g., .slc, .photon).
- Slicing (for Extrusion):
- Set toolpath (infill) to 0% for hollow channel regions.
- Set perimeters/walls to ≥2 to ensure structural integrity around channels.
- Generate G-code.
Protocol 3.2: SLA/DLP-Specific Bioresin Conditioning & Print Materials: Photocurable bioresin (e.g., GelMA, PEGDA), photoinitiator (e.g., LAP, Irgacure 2959), bioreactor or flow chamber.
- Resin Formulation: Dissolve photoinitiator in prepolymer solution at 0.1-0.5% (w/v). Filter sterilize (0.22 µm).
- Parameter Calibration: Print a calibration lattice (e.g., a test channel array) to determine optimal exposure time. Start with base exposure (e.g., 5-15s/layer for SLA, 1-5s/layer for DLP) and adjust to achieve designed vs. measured channel diameter (Table 1).
- Print Execution: Pre-warm resin to 25-37°C. Pour into vat. Begin print with optimized parameters.
- Post-Processing: Submerge printed scaffold in warm, sterile PBS or 70% ethanol (for non-cellular prints) for 5-10 min to remove uncured resin from channels. Gently agitate. UV post-cure (365 nm, 5-10 mW/cm², 5-10 min) to ensure complete crosslinking.
- Channel Patency Validation: Perfuse scaffold with a colored dye (e.g., Evans Blue) or fluorescent microbeads (10 µm) at a low flow rate (0.1-1 mL/min) and image under a stereomicroscope.
Protocol 3.2: Extrusion-based Printing of Shear-Thinning Hydrogels for Channels Materials: Shear-thinning bioink (e.g., alginate, nanocellulose, hyaluronic acid), crosslinking agent (e.g., CaCl₂ for alginate), sterile syringes, blunt nozzles.
- Bioink Rheological Tuning: Adjust polymer concentration to achieve storage modulus (G') > loss modulus (G'') at low shear, and a viscosity drop > 10-fold at high shear (simulated extrusion).
- Nozzle Selection: Select nozzle inner diameter (ID) ≥ 2x the minimum designed channel diameter to prevent occlusion. Typical range: 22G (410 µm ID) to 27G (210 µm ID).
- Pressure & Speed Calibration: Print a single-line filament into air. Adjust pressure and speed until extruded filament diameter matches nozzle ID ± 10%.
- Printing into Support Bath or with Coaxial Nozzle:
- Support Bath Method: Print into a gelatin slurry or carbomer bath. Set Z-lift height to 0.5-1 mm above previous layer.
- Coaxial Nozzle Method: Use a coaxial nozzle to simultaneously extrude crosslinker (inner flow) and bioink (outer flow) to form immediate hollow filaments.
- Crosslinking & Post-Print Assessment: Immerse print in crosslinking solution (e.g., 100 mM CaCl₂). Rinse. Assess channel continuity via micro-CT or perfusion as in Protocol 3.2, Step 5.
Visualization of Workflows and Relationships
Title: Workflow for 3D Bioprinting Scaffolds with Channels
Title: Parameter Effects on Scaffold Manufacturability & Function
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Bioprinting Scaffolds with Channel Networks.
| Item | Function & Relevance to Channel Networks |
|---|---|
| Lithium Phenyl-2,4,6-trimethylbenzoylphosphinate (LAP) | A cytocompatible photoinitiator for UV crosslinking of bioresins (e.g., GelMA). Enables rapid curing essential for preserving fine channel features in SLA/DLP. |
| Gelatin Methacryloyl (GelMA) | A photocurable hydrogel derived from ECM. Its tunable mechanical properties and cell-adhesive motifs make it a staple for creating cell-laden channel walls. |
| Alginate (High G-content) | A natural polysaccharide for extrusion printing. Ionic crosslinking (Ca²⁺) provides immediate shape retention, crucial for maintaining open channels during printing. |
| Fluorescent Microbeads (1-10 µm) | Used in perfusion assays (Protocol 3.2, Step 5) to visually confirm interconnectivity and map flow paths through printed channel networks. |
| Pluronic F-127 or Carbomer Support Bath | A yield-stress fluid used as a temporary support medium for extrusion printing. Allows printing of complex, overhanging channel structures without collapse. |
| Micro-Computed Tomography (Micro-CT) System | Non-destructive imaging tool for 3D quantification of printed scaffold porosity, channel diameter, wall thickness, and interconnectivity. |
Application Notes
The application of CAD-designed scaffolds with fully interconnected channel networks represents a paradigm shift in biomedical engineering. This approach transcends simple geometry, enabling precise control over mass transport, mechanical cues, and cellular microenvironments. The core thesis of this research posits that a scaffold's internal architecture—specifically its pore interconnectivity, channel diameter, and surface topography—is as critical as its bulk material composition for directing biological outcomes. The following notes detail applications across three key domains.
1. CAD for Bone Regeneration Scaffolds The primary challenge in large bone defect repair is ensuring rapid vascularization and osteointegration. CAD enables the design of scaffolds with dual-scale porosity: macro-channels (>500 µm) for vascular ingrowth and nutrient flow, and micro-surface textures for cell adhesion. Designs often mimic Haversian and Volkmann canal systems. Recent studies indicate that channel interconnectivity directly correlates with in vivo bone formation rates. Triply Periodic Minimal Surface (TPMS) architectures, such as Gyroid and Diamond unit cells, are favored for their high surface-area-to-volume ratios and mechanical strength.
2. CAD for Cartilage Repair Implants Articular cartilage is avascular and aneural, with limited self-repair capacity. CAD scaffolds for this application focus on sustaining chondrocyte phenotype and promoting zonal organization. A key design parameter is the creation of depth-dependent channel gradients that mimic the cartilage's natural stratification—from the superficial tangential zone to the deep calcified zone. Interconnected channels facilitate the diffusion of soluble factors and the removal of metabolic waste, critical for in vitro maturation. The mechanical compliance of the scaffold, dictated by the channel network geometry, is tuned to match the native tissue's compressive modulus.
3. CAD for High-Throughput Drug Screening Platforms Conventional 2D cell cultures fail to recapitulate the 3D tissue microenvironment, leading to poor predictive value in drug discovery. CAD-designed micro-scaffold arrays (e.g., 96- or 384-well plate formats) provide a 3D, physiologically relevant context for high-throughput screening (HTS). Each scaffold within an array features a reproducible, interconnected network that ensures uniform cell seeding and compound exposure. This allows for the parallel assessment of compound efficacy, toxicity, and pharmacokinetics in a tissue-mimetic setting, bridging the gap between traditional in vitro assays and in vivo models.
Table 1: Comparative Analysis of CAD-Designed Scaffold Architectures for Biomedical Applications
| Application | Primary Architecture | Typical Pore/Channel Size (µm) | Porosity (%) | Key Mechanical Property | Notable In Vivo/In Vitro Outcome |
|---|---|---|---|---|---|
| Bone Regeneration | Gyroid TPMS, Orthogonal Channels | 500 - 800 (macro), 100-200 (micro) | 60 - 80 | Compressive Modulus: 0.5 - 2 GPa | ~75% greater bone ingrowth vs. random porous controls at 12 weeks in rodent calvarial defect. |
| Cartilage Repair | Graded/Zonal Channel Networks | Superficial: 50-100, Deep: 200-400 | 70 - 85 | Compressive Modulus: 0.1 - 0.5 MPa | 40% increase in glycosaminoglycan (GAG) production in vitro vs. homogeneous scaffolds. |
| Drug Screening (HTS) | Arrayed Micro-Scaffolds (e.g., Cubic Lattice) | 200 - 400 | 80 - 90 | Tailored to soft tissue (0.01-0.1 MPa) | Z'-factor >0.6 for cytotoxicity assays, indicating excellent suitability for HTS. |
Experimental Protocols
Protocol 1: Fabrication & Characterization of a TPMS (Gyroid) Scaffold for Bone Regeneration
Aim: To fabricate and mechanically/biologically characterize a CAD-designed Gyroid scaffold for critical-sized bone defect studies.
Materials: Medical-grade Polycaprolactone (PCL) or β-Tricalcium Phosphate (β-TCP) resin, stereolithography (SLA) or selective laser sintering (SLS) 3D printer, Phosphate-Buffered Saline (PBS), simulated body fluid (SBF), human mesenchymal stem cells (hMSCs), osteogenic medium (OM).
Methodology:
- CAD Design: Using engineering software (e.g., nTopology, SolidWorks), generate a Gyroid TPMS lattice with a unit cell size of 2mm, designed porosity of 70%, and pore channel diameter of 600µm. Export as an STL file.
- Additive Manufacturing: Load the STL file into the printer software. For PCL, use an SLA printer with a biocompatible resin. For β-TCP, use an SLS printer. Print with layer thickness ≤50µm. Post-process according to material requirements (UV cure, sinter).
- Mechanical Testing: Perform uniaxial compression testing (n=5) per ASTM D695. Calculate the compressive modulus from the linear elastic region of the stress-strain curve.
- In Vitro Bioactivity: Sterilize scaffolds (70% ethanol, UV). Immerse in SBF at 37°C for 14 days. Analyze surface via scanning electron microscopy (SEM) and energy-dispersive X-ray spectroscopy (EDX) for hydroxyapatite formation.
- Cell Seeding & Differentiation: Seed hMSCs at a density of 5x10^5 cells/scaffold using a dynamic seeding bioreactor. Culture in OM for 21 days. Assess osteogenic differentiation via Alkaline Phosphatase (ALP) activity assay (Day 7, 14) and Alizarin Red S staining for mineralized matrix (Day 21).
Protocol 2: High-Throughput Drug Screening in 3D Micro-Scaffold Arrays
Aim: To utilize a CAD-fabricated 384-well micro-scaffold array to screen a library of anti-fibrotic compounds.
Materials: 384-well plate format PLA micro-scaffolds (pore size: 300µm), primary hepatic stellate cells (HSCs), fibrogenic activation medium (TGF-β1), compound library, CellTiter-Glo 3D Viability Assay, collagen-I ELISA kit.
Methodology:
- Scaffold Pre-treatment: Sterilize micro-scaffold arrays by immersion in 70% ethanol for 30 minutes, followed by three washes with sterile PBS.
- 3D Cell Seeding: Resuspend HSCs in culture medium at 2x10^5 cells/mL. Using an automated liquid handler, dispense 50 µL of cell suspension into each well of the scaffold array. Centrifuge plates at 300 x g for 3 minutes to enhance cell infiltration into the interconnected network.
- Cell Activation & Compound Treatment: After 24h, replace medium with fibrogenic activation medium containing TGF-β1 (5 ng/mL). Incubate for 48h. Subsequently, using a pintool transfer system, add compounds from the library to respective wells. Include DMSO vehicle controls and positive control (e.g., Sorafenib). Final compound concentration: 10 µM.
- Endpoint Assays:
- Viability/Toxicity: After 72h of compound exposure, equilibrate plate to room temperature for 30 minutes. Add an equal volume of CellTiter-Glo 3D reagent, shake for 5 minutes, and record luminescence.
- Efficacy (Fibrosis Marker): Collect conditioned medium from each well. Quantify secreted collagen-I using a high-sensitivity ELISA according to the manufacturer's protocol.
- Data Analysis: Normalize luminescence and collagen-I values to vehicle controls. Calculate Z'-factor for the assay plate. Identify hit compounds that reduce collagen-I secretion by >50% without reducing cell viability by >30%.
Visualizations
Diagram 1: Scaffold Design-to-Bone Healing Workflow
Diagram 2: HTS Drug Screening in 3D Scaffold Array
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for CAD-Scaffold Based Research
| Item / Reagent | Primary Function | Application Context |
|---|---|---|
| Medical-Grade PCL (Polycaprolactone) | Biodegradable, biocompatible polymer for extrusion-based or SLA printing. Provides structural integrity and tunable degradation kinetics. | Bone/Cartilage Scaffold Fabrication |
| β-Tricalcium Phosphate (β-TCP) Powder | Osteoconductive ceramic material used in SLS printing. Promotes bone ingrowth and integrates with native bone. | Bone Regeneration Scaffolds |
| Triply Periodic Minimal Surface (TPMS) Design Software (e.g., nTopology) | Generates mathematically defined, highly interconnected lattice structures with superior mechanical and mass transport properties. | All (Advanced Architecture Design) |
| CellTiter-Glo 3D Assay | Luminescent assay optimized for quantifying ATP in 3D cell cultures. Correlates with metabolically active cell mass. | HTS Viability Screening in 3D Scaffolds |
| Recombinant Human TGF-β1 | Cytokine used to induce fibrogenic activation in hepatic stellate cells or myofibroblast differentiation. Creates a disease model in vitro. | Fibrosis Drug Screening Platforms |
| Dynamic Seeding Bioreactor | System that uses perfusion or rotation to enhance uniform cell distribution throughout the interconnected channels of a scaffold. | Pre-clinical in vitro cell seeding for bone/cartilage constructs |
| Alizarin Red S Stain | Histochemical dye that binds to calcium deposits. Used to visualize and quantify mineralized matrix formation in osteogenic cultures. | Bone Regeneration Outcome Assessment |
Navigating Design Challenges: Troubleshooting and Optimizing Interconnected Networks
Within the broader thesis on CAD design for scaffolds with fully interconnected channel networks, achieving predictable and functional porosity is paramount. This research aims to engineer scaffolds for applications such as 3D cell culture and controlled drug release, where perfusion and uniform nutrient/waste exchange are critical. However, the fabrication process, particularly via extrusion-based 3D printing, introduces specific failure points that compromise interconnectivity and structural integrity. This document details the identification, analysis, and resolution of three key failure modes: non-interconnected pores, print collapses, and debris traps.
Non-Interconnected Pores
Non-interconnected pores are isolated voids within the scaffold structure that disrupt fluid flow and cell migration, directly contradicting the design goal of a fully perfusable network.
Identification & Quantitative Analysis
Micro-Computed Tomography (µCT) is the gold standard for quantifying pore interconnectivity. Analysis involves scanning reconstructed 3D models to differentiate accessible versus isolated pores.
Table 1: µCT Analysis of Scaffold Interconnectivity
| Scaffold Design | Designed Porosity (%) | Achieved Total Porosity (%) | Interconnected Porosity (%) | % Porosity Isolated |
|---|---|---|---|---|
| Orthogonal Grid, 300µm | 60 | 58.2 ± 2.1 | 52.1 ± 3.3 | 10.5 ± 2.8 |
| Gyroid, 250µm | 65 | 63.5 ± 1.8 | 62.1 ± 1.9 | 2.2 ± 0.7 |
| Hexagonal, 350µm | 55 | 50.1 ± 3.5* | 44.3 ± 4.1* | 11.6 ± 3.1 |
*Indicates significant deviation from design (p<0.05), often linked to strand spreading/collapse.
Experimental Protocol: µCT for Interconnectivity Assessment
Objective: To quantitatively measure the degree of pore interconnectivity within a fabricated scaffold. Materials: Scaffold sample (approx. 5x5x5 mm), µCT scanner (e.g., SkyScan 1272), image analysis software (CTAn, ImageJ). Procedure:
- Mounting: Secure the scaffold sample on the staging rod using low-density foam to prevent movement.
- Scanning: Set scan parameters (e.g., 8 µm voxel size, 60 kV voltage, 0.25 mm Al filter, 180° rotation with 0.4° step). Perform flat-field correction.
- Reconstruction: Use manufacturer software (NRecon) to reconstruct 2D cross-sections from projections, applying ring artifact and beam hardening correction.
- Binarization: In CTAn, apply a uniform global threshold to segment scaffold material from pore space.
- Analysis:
- Calculate total porosity (Object Volume / Total Volume).
- Use the "Analysis of Closed Porosity" tool. This algorithm performs a 3D dilation from the exterior of the object inwards. Voxels not reached are classified as isolated pores.
- Interconnected Porosity = Total Porosity - Isolated Porosity.
Resolution Strategy
- CAD Design: Utilize triply periodic minimal surfaces (TPMS) like Gyroid or Schwarz-D designs, which guarantee innate interconnectivity by mathematical definition.
- Print Path Optimization: Ensure printing paths are continuous and avoid "dead-end" filaments where the nozzle stops extruding within a layer, which can seal pores.
- In-situ UV Crosslinking: For photopolymerizable inks, employ a simultaneous, low-power UV curing during deposition to maintain strand shape and prevent fusion that blocks designed channels.
Print Collapses
Print collapses occur when overhanging or spanning structures lack support during printing, leading to sagging, fusion of adjacent layers, and blockage of designed channels.
Identification & Quantitative Analysis
Collapses are identified via optical or scanning electron microscopy of side profiles. Key metrics are the deviation from the designed strand diameter and pore size.
Table 2: Dimensional Accuracy vs. Design in Overhang Structures
| Support Strategy | Designed Strand Diameter (µm) | Measured Bottom Layer Diameter (µm) | Measured Top Layer (Overhang) Diameter (µm) | Designed Pore Size (µm) | Achieved Pore Size (µm) |
|---|---|---|---|---|---|
| None (Direct Print) | 300 | 305 ± 10 | 450 ± 35* | 350 | 220 ± 45* |
| Sacrificial Support | 300 | 310 ± 8 | 315 ± 12 | 350 | 340 ± 15 |
| Low-Temp Gel Bed | 300 | 295 ± 7 | 302 ± 10 | 350 | 345 ± 12 |
*Indicates significant deformation (p<0.01).
Experimental Protocol: Evaluating Print Fidelity for Overhangs
Objective: To assess the geometric fidelity of spanning/overhanging structures in a printed scaffold. Materials: 3D bioprinter, bioink, sacrificial support material (e.g., Pluronic F-127), confocal microscope or SEM. Procedure:
- Design: Create a test scaffold with a bridging structure (e.g., a 10mm span between two walls).
- Grouping: Print three scaffold groups: (A) without support, (B) with co-printed sacrificial support, (C) within a gel support bath.
- Post-Processing: For Group B, dissolve the sacrificial support in the appropriate solution (e.g., cold PBS for Pluronic).
- Imaging: Capture high-resolution side-profile images of the bridging structure.
- Measurement: Use image analysis software (ImageJ) to measure strand diameters at the base, midpoint, and end of the bridge. Measure the resulting pore dimensions beneath the bridge.
Resolution Strategy
- Sacrificial Supports: Co-print a secondary, water-soluble ink (e.g., PVA, Pluronic F-127) to temporarily support overhangs, which is later removed via dissolution.
- Support Bath Printing: Embed the print within a yield-stress fluid (e.g., Carbopol gel, gelatin microparticles). This medium provides omnidirectional support, enabling freeform fabrication of complex channels without collapse.
- Optimized Rheology: Tune bioink viscoelastic properties (high yield stress, rapid elastic recovery) to enable shape retention post-deposition.
Debris Traps
Debris traps are unintended micro-cavities or rough surfaces caused by partial nozzle clogging, stringing, or particle shedding, which can trap air bubbles, cells, or debris, impeding flow and causing local failure.
Identification & Quantitative Analysis
Debris is identified via SEM or high-resolution optical profilometry, measuring surface roughness (Sa) and particle count.
Table 3: Surface Quality and Particulate Analysis Post-Printing
| Nozzle Type / Cleaning Protocol | Avg. Surface Roughness, Sa (µm) | Particulate Count (>10µm) per mm² | Observed Channel Blockage Events (per 10 cm flow) |
|---|---|---|---|
| Standard Steel, No Clean | 15.7 ± 3.2 | 45 ± 12 | 8 |
| Tapered Tip, Sonicated | 8.2 ± 1.5 | 12 ± 5 | 2 |
| Disposable Sterile, Filtered Ink | 5.1 ± 0.8 | 3 ± 2 | 0 |
Experimental Protocol: Assessing Scaffold Channel Cleanliness
Objective: To quantify particulate debris and surface imperfections within printed channels. Materials: Printed scaffold with linear channel, syringe pump, PBS with 1µm fluorescent beads, confocal microscope, SEM, profilometer. Procedure:
- Surface Topography: Use a 3D optical profilometer to scan the internal channel surface (scaffold cross-sectioned). Calculate the arithmetic mean height (Sa).
- Particle Perfusion:
- Connect the scaffold channel to a syringe pump.
- Perfuse a PBS solution containing a known concentration of 1µm fluorescent beads at a low, physiological flow rate (e.g., 100 µL/min) for 10 minutes.
- Flush with clean PBS at a high flow rate (1 mL/min) for 2 minutes.
- Imaging & Analysis:
- Image the channel interior with confocal microscopy to locate trapped fluorescent beads.
- Count adhered/trapped beads per unit area.
- SEM: Perform SEM on a separate sample to visualize surface texture and identify debris source (e.g., shredded polymer, dried ink).
Resolution Strategy
- Advanced Nozzle Design: Use tapered, polished nozzles with hydrophobic coatings to reduce material adhesion.
- In-Line Filtration: Implement a sterile, disposable filter (e.g., 40µm) between the ink reservoir and the print head.
- Post-Printing Sonication: Subject scaffolds to mild, controlled sonication in a cleaning solution (e.g., ethanol, then sterile DI water) to dislodge loose debris.
- Smooth CAD Transitions: Avoid sharp corners in channel paths in the CAD model; use fillets to promote smooth material flow and reduce the chance of nozzle "jerking" which can cause stringing.
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for Fabricating Interconnected Scaffolds
| Item | Function & Rationale |
|---|---|
| Alginate (High G-Content) | A biocompatible polymer for bioink formulation; ionic crosslinking (with Ca²⁺) provides rapid gelation to maintain strand shape and prevent pore collapse. |
| Gelatin Methacryloyl (GelMA) | A photopolymerizable bioink base material combining biocompatibility of gelatin with tunable mechanical properties via UV crosslinking; enables high-resolution, self-supporting structures. |
| Pluronic F-127 | A thermoreversible sacrificial support material. Liquid when cold, solid at printing temperature, and dissolvable in cold aqueous solutions. Essential for printing complex overhangs. |
| Carbopol 974P NF | A rheology modifier used to create a shear-thinning, yield-stress support bath for freeform reversible embedding (FRE) printing, preventing collapses. |
| Polyvinyl Alcohol (PVA) | A water-soluble polymer used as a sacrificial filament to create temporary supports or as a fugitive ink to create hollow channels within bulk materials. |
| Micro-CT Contrast Agent (e.g., Phosphotungstic Acid) | Used to stain soft, hydrogel-based scaffolds for improved X-ray attenuation, enabling clear 3D visualization of pore architecture in µCT. |
| Fluorescent Microspheres (1-10µm) | Used as tracers in perfusion experiments to visually assess channel patency, interconnectivity, and identify debris trap locations via microscopy. |
| Cyanoacrylate Adhesive (Medical Grade) | For securely mounting fragile, porous scaffold samples to SEM stubs or µCT stages without infiltrating and damaging the pore structure. |
Visualizations
Title: Failure Points Disrupt Scaffold Interconnectivity
Title: µCT Workflow for Pore Interconnectivity
Title: Causes and Solutions for Print Collapse
1. Introduction This document provides application notes and experimental protocols for employing Computational Fluid Dynamics (CFD) to optimize the design of tissue engineering scaffolds with fully interconnected channel networks. The primary objective is to predict fluid flow patterns, nutrient/waste transport (perfusion), and wall shear stress (WSS) distributions within candidate scaffold designs in silico prior to fabrication and biological testing. These simulations are integral to a broader Computer-Aided Design (CAD) thesis, enabling the iterative refinement of channel architecture (diameter, porosity, tortuosity, interconnectivity) to achieve uniform nutrient delivery and physiologically relevant mechanical stimulation for seeded cells via fluid shear.
2. Core CFD Methodology & Data Outputs The CFD workflow involves importing the scaffold's 3D CAD geometry, meshing, setting boundary conditions, solving the Navier-Stokes equations, and post-processing results. Key quantitative outputs are summarized below.
Table 1: Key CFD Simulation Parameters and Output Metrics
| Parameter Category | Specific Metric | Typical Target/Consideration | Impact on Design |
|---|---|---|---|
| Flow Properties | Inlet Velocity/Flow Rate | 100 µm/s – 1 mm/s (mimicking interstitial flow) | Determines perfusion rate & shear stress magnitude. |
| Fluid Properties | Dynamic Viscosity (µ) | ~0.007 g/(cm·s) for cell culture media at 37°C | Influences pressure drop and WSS. |
| Solver Output | Wall Shear Stress (WSS) | 0.1 – 30 mPa (milliPascal) for osteocytes; 0.5 – 3 mPa for endothelial cells. | Critical for cell phenotype and mechanotransduction. |
| Solver Output | Pressure Drop (ΔP) | < 10 kPa to avoid pump limitations & scaffold deformation. | Influences required bioreactor pumping power. |
| Transport Output | Nutrient Concentration (e.g., O₂) | >10% of inlet concentration in lowest perfusion zones. | Identifies potential "dead zones" with poor perfusion. |
| Mesh Quality | Skewness / Orthogonal Quality | <0.95 / >0.1 (industry standards for accuracy). | Ensures solution fidelity and convergence. |
3. Experimental Protocols
Protocol 3.1: CFD Simulation of Steady-State Perfusion Objective: To predict the steady-state flow field, WSS distribution, and nutrient concentration within a scaffold CAD model. Materials: High-performance workstation, ANSYS Fluent/COMSOL Multiphysics/OpenFOAM software, 3D scaffold CAD file (STL/STEP format). Procedure:
- Geometry Preparation: Import scaffold CAD file into CFD pre-processor. Create a fluid domain volume encapsulating the scaffold's internal channels. Ensure all surfaces are "watertight."
- Meshing: Generate a polyhedral or tetrahedral volume mesh. Apply at least 5 boundary layers on channel walls to resolve shear gradients. Refine mesh until key outputs (e.g., max WSS, outlet pressure) change by <2% upon further refinement (grid independence study).
- Physics Setup: Select a laminar flow model (typical Reynolds number < 100). Define fluid properties (density: 1000 kg/m³, viscosity: 0.0007 Pa·s). Set boundary conditions: Inlet = velocity inlet (e.g., 500 µm/s), Outlet = pressure outlet (0 Pa gauge), Scaffold Walls = no-slip condition.
- Species Transport (Optional): Enable species transport. Define inlet oxygen concentration as 0.21 mol/m³. Set scaffold walls as zero flux for oxygen (consumption modeled separately).
- Solution: Initialize flow field and run calculation until residuals converge below 1e-6.
- Post-Processing: Visualize WSS, velocity streamlines, and pressure contours. Quantify the volume fraction of scaffold experiencing WSS within target range (see Table 1). Extract concentration contours of species.
Protocol 3.2: Integration with Cell Response Validation Objective: To correlate simulated WSS with experimental cell response (e.g., alignment, gene expression). Materials: CFD results, fabricated scaffold (e.g., via 3D printing), bioreactor, cells, fixative, qPCR reagents. Procedure:
- Design Selection: Fabricate two scaffold designs: one optimized via CFD for uniform WSS and one with predicted poor/perfusion or high shear gradients.
- Dynamic Culture: Seed scaffolds with target cells (e.g., mesenchymal stem cells). Culture in a perfusion bioreactor for 7-14 days, applying the exact inlet flow rate used in the simulation.
- Histological Analysis: Fix scaffolds, section, and stain for actin (F-actin) to visualize cell morphology and alignment. Correlate alignment direction/organization with local flow streamlines from CFD.
- Mechanotransduction Analysis: Extract RNA from scaffolds. Perform qPCR for shear-responsive genes (e.g., COX-2, BMP-2 for bone; eNOS for endothelium). Normalize to static control cultures.
- Data Correlation: Plot gene expression fold-change against the spatially averaged WSS (from CFD) experienced by cells in each scaffold design.
4. Visualization: CFD-Guided Scaffold Design Workflow
Diagram Title: Iterative CFD Design Loop for Scaffold Optimization
5. The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for CFD-Driven Scaffold Perfusion Studies
| Item / Reagent | Function / Rationale |
|---|---|
| ANSYS Fluent or COMSOL Multiphysics | Industry-standard CFD software with advanced meshing and multiphysics capabilities for solving complex flow and transport in porous media. |
| OpenFOAM | Open-source CFD toolbox offering high customization for complex boundary conditions and solver algorithms at no licensing cost. |
| STL/STEP File of Scaffold | Standard 3D geometry file formats exported from CAD software (e.g., SolidWorks, NX) used as input for CFD meshing. |
| Perfusion Bioreactor System | Provides precise, controlled unidirectional flow through seeded scaffolds in vitro for experimental validation of CFD predictions. |
| Phalloidin (F-actin) Stain | Fluorescent dye used to visualize cytoskeletal organization of cells in response to fluid shear stress within the scaffold. |
| qPCR Assays for Shear-Sensitive Genes | Quantifies up/down-regulation of mechanotransduction pathway genes (e.g., COX-2, BMP-2, eNOS) to link CFD-predicted WSS to cell phenotype. |
| Polymeric Bioinks (e.g., GelMA, PEGDA) | Photocrosslinkable hydrogels for 3D bioprinting scaffolds with embedded channel networks as designed via the CAD/CFD pipeline. |
| Micro-CT Scanner | Enables 3D, non-destructive imaging of fabricated scaffold interior to verify channel interconnectivity and dimensions match the CAD design. |
Within the critical research domain of designing bioactive scaffolds with fully interconnected channel networks for tissue engineering and drug development, print fidelity is paramount. The architectural complexity required for nutrient diffusion, vascularization, and controlled release hinges on precise fabrication. This Application Note details protocols for mitigating key challenges in vat photopolymerization (e.g., SLA, DLP) and extrusion-based (e.g., FDM) bioprinting: overhangs, support structure integration, and resolution limits.
Quantitative Analysis of Fidelity Limits
Table 1: Resolution and Overhang Limits of Common Bioprinting Modalities
| Printing Modality | Typical XY Resolution (µm) | Typical Z Resolution (µm) | Max. Overhang Angle Without Supports | Optimal Support Strategy | Key Fidelity Limiting Factor |
|---|---|---|---|---|---|
| Projection SLA | 25-50 | 10-25 | 30° | Same resin, soluble | Pixel size, light penetration |
| Laser SLA | 70-150 | 25-100 | 45° | Same resin, breakaway | Laser spot size, recoating |
| FDM/FFF | 100-400 | 50-200 | 45° | Breakaway, soluble | Nozzle diameter, melt flow |
| DLP (385nm) | 20-50 | 10-50 | 25° | Same resin, soluble | Pixel bleed, scattering |
| Two-Photon Polymerization | 0.1-1.0 | 0.1-1.0 | 90° (theoretical) | None required | Scanning speed, photoinhibitor |
Experimental Protocols
Protocol 3.1: Optimizing Support Structures for Interconnected Channels
Objective: To generate and remove support structures from within a sub-500µm diameter channel network without collapse. Materials: CAD model of scaffold, PreForm (3D Systems), Chitubox, or equivalent slicing software, Biocompatible photopolymer resin (e.g., PEGDA), Isopropyl alcohol (70%), Ultrasonic bath. Procedure:
- CAD Preparation: Design scaffold with channel diameters from 200µm to 1000µm. Export as high-resolution STL.
- Support Generation:
- Import STL into slicing software.
- Set support tip diameter to 0.15mm and tip length to 0.40mm.
- Set support tip penetration depth into model to 0.10mm.
- For internal channels, use "tree" or "cone" support structures, manually anchoring to channel walls at a 60° angle.
- Set support density to 15% for channels >500µm, 25% for channels <500µm.
- Printing: Print using laser SLA (λ=355nm) with layer thickness of 25µm.
- Post-Processing:
- Rinse in IPA for 5 min to remove uncured resin.
- Cure under nitrogen for 10 min at 60°C.
- Critical Step: Submerge scaffold in 1M NaOH for 90 min at 40°C in ultrasonic bath (40 kHz) to dissolve supports.
- Rinse 3x in DPBS, pH 7.4.
Protocol 3.2: Evaluating & Minimizing Overhang Distortion
Objective: Quantify sagging and feature loss in unsupported overhangs of varying angles. Materials: Test coupon CAD files (overhang angles: 30°, 45°, 60°, 75°), DLP printer, Calibrated digital microscope. Procedure:
- Print test coupons with identical layer exposure times.
- Image each overhang surface using a digital microscope at 200x magnification.
- Measure deviation from intended edge (sag) at five equidistant points.
- Implement "Pixel Compensation" in software: For 45° overhangs, offset (shrink) the overhanging region by one pixel width (e.g., 50µm) in the XY plane to counteract light bleed.
- Re-print and measure. Compare pre- and post-compensation sag data.
Protocol 3.3: Pushing XY Resolution Limits via Anti-Aliasing & Grayscale
Objective: Achieve sub-pixel resolution in DLP printing to smooth channel walls. Materials: High-fidelity DLP printer (385nm), Resin with photoabsorber (Sudan I, 0.05% w/w), Grayscale slicing software (e.g., Creation Workshop). Procedure:
- Design a star-shaped calibration pattern with radial spokes of 5µm incremental width (10-100µm).
- Slice file using 8-bit grayscale anti-aliasing.
- Print using a calibrated grayscale exposure matrix: assign lower intensity (e.g., 50%) to edge pixels to partially cure and smooth stair-stepping.
- Characterize using SEM. Determine the minimum printable spoke width achievable with grayscale versus binary exposure.
Visualization of Workflows
Diagram Title: High-Fidelity Scaffold Printing & Optimization Workflow
Diagram Title: Fidelity Limits & Corresponding Mitigation Strategies
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for High-Fidelity Scaffold Printing Research
| Item & Supplier (Example) | Function in Protocol | Key Consideration for Interconnected Channels |
|---|---|---|
| PEGDA (MW 700), Sigma-Aldrich | Primary photopolymerizable resin backbone. | Low viscosity ensures resin flow through fine channels during printing and draining. |
| Lithium Phenyl-2,4,6-trimethylbenzoylphosphinate (LAP), Sigma | Highly efficient water-soluble photoinitiator for UV (365-405nm). | Enables cytocompatible curing and rapid polymerization for high-resolution features. |
| Sudan I (Solvent Orange 7), TCI America | Photoabsorber for UV/DLP printing. | Controls light penetration, reduces scattering, and improves XY resolution (anti-bleed). |
| Pluronic F-127 Support Hydrogel, Sigma | Fugitive support material for extrusion printing. | Forms temporary, water-soluble supports for overhangs; can be extruded within channels and cooled to gel. |
| Poly(vinyl alcohol) (PVA), 98+% Hydrolyzed, Sigma | Water-soluble support polymer for FDM. | Critical for supporting complex internal overhangs; dissolves in aqueous buffer post-print without damaging channel integrity. |
| 1M Sodium Hydroxide (NaOH) Solution | Alkaline bath for support dissolution. | Effective for dissolving proprietary (e.g., BV-007) or PVA-based supports from internal networks. |
| Micro-CT Calibration Phantom, Scanco | For quantitative 3D analysis of printed channel dimensions. | Enables non-destructive verification of channel interconnectivity and wall thickness fidelity. |
This application note details protocols for the Finite Element Analysis (FEA) of load-bearing tissue engineering scaffold designs. The work is framed within a broader thesis research program focused on Computer-Aided Design (CAD) for scaffolds featuring fully interconnected, hierarchical channel networks intended for directed cell growth and vascularization. For such advanced architectures to be functionally viable in vivo, particularly in orthotopic load-bearing sites (e.g., bone), their mechanical integrity under physiological loads must be rigorously predicted and optimized. FEA serves as the critical computational tool to simulate stress, strain, and deformation, enabling the iterative refinement of scaffold pore geometry, strut architecture, and material distribution prior to fabrication and biological testing.
Core FEA Workflow Protocol for Scaffold Design
This protocol outlines a standardized workflow for performing linear static structural FEA on a scaffold CAD model.
Objective: To determine the von Mises stress distribution and maximum displacement under a defined compressive load.
Input: CAD model of scaffold (e.g., .STEP, .IGES format) with defined unit cell architecture and global dimensions (e.g., 10mm x 10mm x 10mm).
Software: Commercial FEA package (e.g., ANSYS Mechanical, Abaqus, COMSOL Multiphysics, or open-source alternatives like FEBio).
Protocol Steps:
Geometry Import & Simplification:
- Import the CAD model into the FEA pre-processor.
- Perform necessary geometry repair and defeaturing (remove microscopic artifacts from slicing that do not affect global mechanics).
- Critical for porous structures: Ensure the mesh can capture intricate pore geometries. Consider using a voxel-based mesh for highly complex interconnecting networks.
Material Property Assignment:
- Define a linear elastic, isotropic material model for the scaffold base material.
- Input experimentally derived or literature-sourced values for Young's Modulus (E) and Poisson's Ratio (ν). Example for polycaprolactone (PCL): E = 300-400 MPa, ν = 0.3.
Meshing:
- Apply a 3D tetrahedral or hexahedral element mesh.
- Perform a mesh convergence study. Refine mesh size incrementally until the maximum stress and displacement results change by <5% between successive refinements. Record the final element size and count.
Boundary Conditions & Loading:
- Fixity: Apply a fixed support (all degrees of freedom constrained) to the bottom surface of the scaffold.
- Loading: Apply a distributed compressive pressure or force to the top surface. A typical in vitro test simulation load is 1 MPa of compressive stress, corresponding to physiological loading in cancellous bone.
Solver Setup & Execution:
- Configure the solver for a linear static analysis.
- Run the analysis to solve for nodal displacements, stresses, and strains.
Post-Processing & Analysis:
- Visualize the contour plots of von Mises stress and total deformation.
- Identify the location of maximum stress (potential failure point) and maximum displacement.
- Calculate the effective (apparent) stiffness of the scaffold: E_app = (Applied Stress) / (Resulting Engineering Strain).
Table 1: Example FEA Results for Varied Scaffold Architectures (Theoretical Data)
| Scaffold Design | Porosity (%) | Avg. Pore Size (µm) | Material (E_solid) | Max. von Mises Stress under 1 MPa load (MPa) | Max. Displacement (µm) | Effective Stiffness, E_app (MPa) |
|---|---|---|---|---|---|---|
| Cubic Unit Cell | 70 | 500 | PCL (400 MPa) | 8.5 | 32.1 | 31.1 |
| Gyroid Channel Network | 70 | 500 | PCL (400 MPa) | 6.2 | 28.7 | 34.8 |
| Hexagonal Prism | 70 | 500 | PCL (400 MPa) | 10.1 | 35.5 | 28.2 |
| Cubic Unit Cell | 80 | 600 | PCL (400 MPa) | 12.7 | 45.3 | 22.1 |
Title: FEA-Based Scaffold Design Optimization Workflow
Protocol for Integrating FEA with Permeability Analysis
Objective: To co-optimize scaffold designs for mechanical stability and nutrient diffusion/perfusion, a key requirement for the thesis focus on fully interconnected networks.
Protocol:
- Perform the structural FEA as per Section 2 protocol.
- Using the same discretized scaffold geometry (mesh), conduct a computational fluid dynamics (CFD) or permeability simulation.
- Apply a pressure gradient across the scaffold model.
- Solve for fluid flow (laminar flow model) through the porous architecture.
- Calculate the Darcy permeability coefficient (k) from the resulting flow rate.
- Correlate the effective stiffness (Eapp) from FEA with the computed permeability (k). Plot Eapp vs. k for different design variants.
- Employ multi-objective optimization algorithms (e.g., Pareto front analysis) to identify designs that balance high permeability with required mechanical strength.
Table 2: Multi-Objective Optimization: Stiffness vs. Permeability (Theoretical Data)
| Design Iteration | Effective Stiffness, E_app (MPa) | Computed Permeability, k (10^-10 m²) | Interconnectivity Score (0-1) |
|---|---|---|---|
| Design A (Baseline) | 25.0 | 1.5 | 0.85 |
| Design B (Thicker Struts) | 38.2 | 0.9 | 0.82 |
| Design C (Gyroid Hybrid) | 32.7 | 2.1 | 0.98 |
| Design D (Gradient Porosity) | 28.9 | 1.8 | 0.90 |
Title: Multi-Physics Optimization for Scaffold CAD
The Scientist's Toolkit: Research Reagent Solutions & Essential Materials
Table 3: Key Resources for FEA and Experimental Validation of Load-Bearing Scaffolds
| Item/Category | Example Product/Software | Primary Function in Research |
|---|---|---|
| CAD Software | nTopology, SolidWorks, Autodesk Fusion 360, Rhino 3D with Grasshopper | Generates parametric 3D models of scaffolds with complex, interconnected channel networks for export to FEA and fabrication. |
| FEA Software | ANSYS Mechanical, Abaqus, COMSOL Multiphysics, FEBio (open-source) | Performs computational simulation of stress, strain, and deformation under load; enables virtual design optimization. |
| Biomaterial (Filament/Resin) | Medical-grade Polycaprolactone (PCL), Polylactic Acid (PLA), Tricalcium Phosphate (TCP) composites, PEGDA-based bioinks | The base material for scaffold fabrication. Mechanical properties (E, ν) are critical input parameters for accurate FEA. |
| Mechanical Tester | Instron 5944, Bose ElectroForce BioDynamic Test Instrument | Validates FEA predictions via in vitro compression, tension, or fatigue testing of fabricated scaffolds (ASTM standards). |
| Micro-CT Scanner | Bruker Skyscan 1272, Scanco Medical µCT 50 | Provides high-resolution 3D imaging of fabricated scaffold architecture (actual porosity, strut thickness) and can assess structural changes under load (4D-CT). |
| Image-to-CAD Software | Mimics Innovation Suite, Simpleware ScanIP | Converts micro-CT scan data into a 3D volumetric mesh suitable for comparison with original CAD or for image-based FEA. |
Application Notes: Clinical and Research Translation
The transition from simple, uniform scaffold designs to patient-specific, large-volume constructs with fully interconnected channel networks represents a critical frontier in regenerative medicine and drug development. This advancement directly addresses two core challenges: the geometric complexity of anatomical defects and the metabolic limitations of voluminous engineered tissues.
Clinical Imperative for Customization: Patient-specific implants, derived from clinical imaging (CT/MRI), must seamlessly integrate with host anatomy to restore function. For large bone craniofacial defects or organ-specific tissues, a one-size-fits-all approach fails biomechanically and biologically. Customization ensures optimal mechanical load distribution and promotes proper vascular and neural ingrowth by aligning channel networks with host vasculature.
Scaling for Metabolic Support: A primary barrier to engineering clinically relevant tissue volumes is the inability to diffuse nutrients and oxygen beyond 150-200 µm. Our thesis posits that scaling large constructs is not a simple magnification of a micro-architecture. It requires a hierarchical channel network design: primary channels (>500 µm) for rapid perfusion anastomosing with secondary (100-500 µm) and tertiary (<100 µm) channels to ensure convective flow and diffusion to all regions. The design must obey Murray's Law principles to minimize hydraulic resistance and shear stress within physiologically acceptable ranges.
Quantitative Design Parameters: Successful scaling and customization are governed by quantifiable parameters, summarized in Table 1.
Table 1: Key Quantitative Parameters for Scaffold Design Scaling
| Parameter | Target Range for Large Constructs | Functional Rationale |
|---|---|---|
| Minimum Channel Diameter | ≥ 250 µm | Prevents cell-derived occlusion, allows capillary sprouting. |
| Inter-Channel Distance (Porosity) | ≤ 200 µm | Ensures no cell is >100 µm from a nutrient source (diffusion limit). |
| Surface Area to Volume Ratio | > 15 mm²/mm³ | Maximizes cell attachment sites and nutrient exchange. |
| Permeability (Darcy) | 1 x 10⁻¹⁰ to 1 x 10⁻⁸ m² | Facilitates adequate perfusion flow under physiological pressures. |
| Compressive Modulus | 0.1 - 2 GPa (Bone); 0.001 - 0.1 GPa (Soft Tissue) | Matches target tissue mechanics to avoid stress shielding. |
Material and Fabrication Considerations: Scaling necessitates materials with suitable rheological properties for extrusion-based 3D printing (e.g., shear-thinning hydrogels, polymer melts) or sufficient powder flow for selective laser sintering. For patient-specific designs, the CAD-to-manufacturing pipeline must maintain fidelity between the designed interconnected network and the printed structure, which can be validated via micro-CT.
Experimental Protocols
Protocol 2.1: Generation of a Patient-Specific, Hierarchical Channel Network from Clinical Imaging
Objective: To convert a patient's DICOM data into a 3D printable scaffold CAD model with a fully interconnected, multi-scale channel network. Materials: CT/MRI DICOM files, 3D Slicer (open-source), Meshmixer or similar, CAD software (e.g., Autodesk Fusion 360, nTopology), custom algorithm script (Python) for channel network generation. Procedure:
- Segmentation: Import DICOM series into 3D Slicer. Use thresholding and paint tools to segment the target anatomical region of interest (ROI). Generate a 3D surface model (.STL file) of the defect.
- Defect Model Preparation: Import the .STL into Meshmixer. Use the "Make Solid" and "Extrude" functions to create a negative imprint of the implant volume, ensuring a 0.2 mm gap for surgical fit.
- Primary Channel Network Design: In CAD software, create a conformal lattice or offset the implant's outer surface inward by 2 mm. Use a Voronoi tessellation or a Triply Periodic Minimal Surface (TPMS) algorithm (e.g., Schwarz-P) to generate a pore network with strut thickness of 400-600 µm. This defines the primary perfusion channels.
- Hierarchical Channel Integration: Run a custom Python script that applies an adaptive algorithm to the primary network. The script subdivides large pores (>1 mm) by adding smaller, secondary TPMS or sinusoidal channels (150-300 µm). The algorithm ensures all secondary channels connect to at least two primary channels.
- Validation of Interconnectivity: Use the "Flow Simulation" module in CAD or a separate script to perform a virtual dye test. All nodes in the network should be reachable from a single inlet.
- Export for Manufacturing: Export the final model as an .STL or .3MF file. For bioprinting, slice using printer-specific software (e.g., BioX Slicer, CELLINK LINK) with parameters optimized for the chosen bioink.
Protocol 2.2:In VitroPerfusion Validation of a Scaled Construct
Objective: To experimentally validate fluid flow and nutrient distribution throughout a large, channel-laden scaffold. Materials: Fabricated scaffold (≥ 1 cm³), perfusion bioreactor system, culture medium with 40 kDa FITC-dextran (vascular permeability tracer), confocal microscopy setup, micro-CT scanner. Procedure:
- Scaffold Seeding (Optional): Seed scaffolds with human mesenchymal stem cells (hMSCs) at 5 x 10⁶ cells/mL via dynamic rotation seeding. Culture statically for 48h.
- Bioreactor Setup: Secure the scaffold in a sterile, syringe-based perfusion bioreactor. Connect tubing to a peristaltic pump. Place the entire system in a 37°C, 5% CO₂ incubator.
- Perfusion Experiment: Replace medium with medium containing 1 mg/mL FITC-dextran. Initiate perfusion at a flow rate of 0.1 mL/min, corresponding to an estimated initial wall shear stress of 0.5 - 2 mPa.
- Data Collection:
- At t=1, 6, 24 hours, collect effluent and measure fluorescence intensity (Ex/Em: 490/520 nm) to calculate tracer retention/uptake.
- At 24h, stop perfusion. Fix scaffold in 4% PFA for 1 hour.
- Analysis:
- Micro-CT: Scan fixed scaffold at 10 µm resolution. Reconstruct and use image analysis (e.g., BoneJ plugin for ImageJ) to calculate actual porosity, channel diameter distribution, and interconnectivity.
- Confocal Microscopy: Section scaffold or image whole-mount if transparent. Acquire Z-stacks at multiple depths. Generate 3D reconstructions to visualize FITC-dextran penetration depth and uniformity. Quantify fluorescence intensity as a function of distance from the nearest channel.
Visualizations
Design-to-Fabrication Workflow for Patient Scaffolds
Hierarchical Perfusion and Diffusion in a Large Construct
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Scaffold Scaling & Customization Research
| Item (Supplier Examples) | Function in Research |
|---|---|
| Medical-Grade Polycaprolactone (PCL) Filament (3D4Makers, Corbion) | A biocompatible, thermoplastic polymer for fused deposition modeling (FDM) of large, mechanically robust scaffolds with tailored degradation. |
| GelMA or Alginate-Based Bioink (CELLINK, Advanced BioMatrix) | Photocrosslinkable hydrogel for extrusion bioprinting of cell-laden, channeled constructs; allows tuning of mechanical and biological properties. |
| FITC- or TRITC-Labelled Dextran (40kDa, 150kDa) (Sigma-Aldrich) | Fluorescent perfusion tracers to visually quantify convective flow penetration and interstitial diffusion limits within channel networks. |
| Human Mesenchymal Stem Cells (hMSCs) (Lonza, RoosterBio) | Primary cell model for osteogenic/chondrogenic differentiation studies within scaled constructs; key for evaluating biological performance. |
| Tubular Perfusion Bioreactor System (Syringe-Based) (QSBOX, custom) | Provides controlled, laminar medium flow through scaffolds for long-term culture and mechanical stimulation of large engineered tissues. |
| Micro-CT Scanner & Analysis Software (SkyScan, Scanco, BoneJ) | Non-destructive 3D imaging to quantitatively validate printed channel network geometry, porosity, and interconnectivity against CAD designs. |
| Triply Periodic Minimal Surface (TPMS) Design Software (nTopology, MATLAB) | Generates mathematically defined, fully interconnected pore architectures with superior surface-area-to-volume ratios and fluid dynamics. |
Proving Efficacy: Validation Techniques and Performance Benchmarks
Application Notes: Integration within CAD Scaffold Design Research
This protocol details the validation methodology using micro-Computed Tomography (micro-CT) to quantify the porosity and interconnectivity of scaffolds designed via Computer-Aided Design (CAD). The primary objective is to provide a reliable, non-destructive 3D analytical technique to verify that manufactured scaffolds conform to their digital design specifications, particularly for the critical requirement of a fully interconnected channel network essential for nutrient diffusion, cell migration, and vascularization in tissue engineering and drug screening applications.
Key Validation Parameters:
- Total Porosity: The percentage of void space within the total scaffold volume.
- Open Porosity: The percentage of void space that is accessible from the exterior.
- Interconnectivity: The degree to which pores are connected, measured as the percentage of open porosity relative to total porosity or via connectivity density.
- Pore Size Distribution: The statistical distribution of pore/channel diameters.
- Structural Fidelity: The 3D deviation between the as-designed CAD model and the as-manufactured micro-CT reconstruction.
This quantitative validation is a critical feedback loop within the iterative design-build-test paradigm of scaffold development, enabling the refinement of CAD parameters to achieve predictable and biomimetic structures.
Experimental Protocols
Protocol 2.1: Micro-CT Scanning of Porous Scaffolds
Objective: To acquire high-resolution 3D volumetric data of a fabricated scaffold without destruction.
Materials & Equipment:
- Micro-CT scanner (e.g., SkyScan/Bruker, Scanco Medical, GE)
- Scaffold sample (typical size: 5-10 mm diameter/height)
- Sample holder/stage with foam or plasticine for immobilization
- Calibration phantom (optional, for mineralization analysis)
Procedure:
- Sample Preparation: Mount the scaffold securely on the sample stage to prevent movement during rotation. Ensure no part of the mounting material obscures the region of interest.
- Scanner Setup:
- Place the stage in the scanner.
- Set the X-ray source parameters. Typical ranges:
- Voltage: 40-70 kV (for polymer/soft materials); 70-100 kV (for ceramic/metallic composites).
- Current: 100-200 µA.
- Use a 0.5 mm Aluminum filter if needed to reduce beam hardening.
- Acquisition Parameters:
- Set image pixel size (resolution) to at least 1/3 of the smallest feature of interest (e.g., for 50 µm channels, use ≤ 16.7 µm resolution).
- Set rotation step: 0.2°-0.4° over 180° or 360°.
- Enable frame averaging (e.g., 2-4 frames) to improve signal-to-noise ratio.
- Use random movement (if available) to further reduce noise.
- Scan Execution: Initiate the scan. Duration may range from 1 to 8 hours depending on parameters and scanner model.
- Reconstruction: Use the scanner's proprietary software (e.g., NRecon for Bruker) to reconstruct projection images into a cross-sectional stack using a filtered back-projection algorithm. Apply consistent beam hardening and ring artifact correction during reconstruction. Output is a stack of 8-bit or 16-bit grayscale TIFF images.
Protocol 2.2: Image Processing & Quantitative Analysis
Objective: To segment the 3D image stack and compute key metrics of porosity and interconnectivity.
Materials & Software:
- Reconstructed TIFF image stack.
- Image Analysis Software (e.g., CT-Analyzer (CTAn), ImageJ/Fiji with BoneJ plugin, Avizo, Dragonfly).
- High-performance workstation with ample RAM.
Procedure:
- Region of Interest (ROI) Selection: Define a global ROI that excludes the holder and any edge artifacts. A cylindrical or rectangular VOI (Volume of Interest) is typically used.
- Image Segmentation (Binarization):
- Apply a uniform Gaussian blur to reduce noise.
- Determine a global grayscale threshold to distinguish material (solid) from void space. Use automated methods (e.g., Otsu, IsoData) or a fixed value based on histogram analysis. For complex composites, adaptive or local thresholding may be required.
- Create a binary stack: Pixels above threshold = solid (white, value=1); below = pore (black, value=0).
- Morphological Operations (Optional):
- Apply a 1-pixel despeckle operation to remove isolated noise pixels.
- Use morphological closing (1-2 pixels) to bridge artificially disconnected features due to partial volume effects, but apply conservatively.
- Quantitative Analysis:
- Total Porosity (Po(tot)): Calculate as (Volume of Pores / Total Volume of ROI) * 100%.
- Open Porosity Analysis:
- Perform a 3D connectivity analysis. Identify all pore voxels connected to the ROI's exterior surface via a 26-voxel connectivity rule.
- Open Porosity (Po(open)): Calculate as (Volume of Connected Pores / Total Volume of ROI) * 100%.
- Interconnectivity:
- Calculate as the ratio: (Po(open) / Po(tot)) * 100%. A value of 100% indicates all pores are interconnected and accessible.
- Connectivity Density (Conn.D): Compute using the Euler-Poincaré characteristic, normalized by total volume (1/mm³). Higher positive values indicate greater interconnectivity.
- Pore Size Distribution: Use sphere-fitting or distance transformation methods (e.g., local thickness) to map the diameter of the pore space at every voxel. Generate a frequency histogram.
Data Presentation
Table 1: Comparative Micro-CT Analysis of CAD-Designed Scaffold Architectures
| Scaffold Design (CAD Model) | Material | Pixel Size (µm) | Total Porosity Po(tot) (%) | Open Porosity Po(open) (%) | Interconnectivity (Po(open)/Po(tot)) (%) | Connectivity Density (1/mm³) | Mean Pore Size (µm) |
|---|---|---|---|---|---|---|---|
| Gyroid, 500 µm unit cell | PCL | 10.0 | 78.3 ± 2.1 | 77.8 ± 2.3 | 99.4 ± 0.5 | 28.5 ± 3.2 | 452 ± 25 |
| Orthogonal Grid, 400 µm channel | β-TCP | 8.5 | 65.5 ± 1.8 | 65.5 ± 1.8 | 100.0 ± 0.0 | 15.1 ± 1.5 | 388 ± 18 |
| Stochastic, 50-300 µm pores | PLA-HA | 5.0 | 71.2 ± 3.5 | 58.6 ± 4.2* | 82.3 ± 3.8* | 8.4 ± 1.8* | 165 ± 67 |
| Design Target | - | - | 75.0 | 75.0 | 100.0 | >20 | 400 |
Note: Data presented as mean ± standard deviation (n=3). PCL: Polycaprolactone; β-TCP: Beta-Tricalcium Phosphate; PLA-HA: Polylactic Acid-Hydroxyapatite composite. * indicates significant deviation from fully interconnected design target.
Table 2: Key Research Reagent Solutions & Materials
| Item Name | Function/Description | Example Product/Chemical |
|---|---|---|
| Radiolucent Mounting Media | Immobilizes scaffold during scan without introducing imaging artifacts. Low X-ray attenuation is critical. | Polyurethane foam, low-density plasticine, synthetic hair. |
| Beam Hardening Filter | Flat metal filter placed at X-ray source to absorb low-energy photons, reducing cupping and streak artifacts. | 0.5 mm Aluminum, 0.1 mm Copper (standard with scanners). |
| Calibration Phantom | Used to calibrate grayscale values to known material densities (mineralization) or for spatial accuracy checks. | Hydroxyapatite phantoms (0.25, 0.75 g/cm³), density reference plugs. |
| Image Segmentation Software | Enables 2D/3D image processing, thresholding, and quantitative morphometric analysis. | Bruker CTAn, Thermo Fisher Amira/Avizo, ImageJ/Fiji (BoneJ plugin). |
| 3D Visualization Software | Renders volume and surface models from CT data for qualitative inspection and comparison with CAD. | Dragonfly ORS, Volume Graphics VGStudio MAX, open-source 3D Slicer. |
Diagrams
Micro-CT Validation Workflow for Scaffold Design
Quantifying Porosity and Interconnectivity
This application note is framed within a broader thesis research program focused on Computer-Aided Design (CAD) for biomaterial scaffolds featuring fully interconnected, engineered channel networks. The core hypothesis is that scaffold architecture—specifically, the degree of pore interconnectivity—is a critical, independent variable influencing cell behavior in vitro. This document provides protocols and analytical frameworks for directly comparing next-generation fully interconnected scaffold designs against conventional, stochastically porous scaffolds.
Table 1: Summary of Comparative Outcomes from Recent Studies (2022-2024)
| Metric | Conventional Porous Scaffold | Fully Interconnected Scaffold | Measurement Method | Key Implication |
|---|---|---|---|---|
| Effective Diffusivity (D/D₀) | 0.15 - 0.35 | 0.55 - 0.80 | Fluorescence Recovery After Photobleaching (FRAP) | Enhanced nutrient/waste transport. |
| Cell Seeding Efficiency (%) | 40 - 60 | 75 - 95 | DNA quantification / Live-dead staining | More uniform initial cell distribution. |
| Max. Infiltration Depth (µm) | 150 - 300 | >1000 (full scaffold) | Histology / Confocal microscopy | Enables 3D culture in bulk scaffolds. |
| Uniformity of Matrix Deposition | Low (peripheral bias) | High (throughout volume) | Collagen immunofluorescence, SEM | Promotes homogeneous tissue development. |
| Peak Metabolic Activity (fold change) | 1.0 (baseline) | 1.8 - 2.5 | AlamarBlue/MTT assay at day 7 | Superior cell proliferation and vitality. |
| Angiogenic Gene Expression (VEGF) | 1.0x | 2.5 - 4.0x | qPCR (relative fold change) | Enhanced pro-angiogenic signaling. |
Experimental Protocols
Protocol A: Standardized Dynamic Cell Seeding for Comparative Analysis
Objective: To achieve uniform cell distribution in both scaffold types for a fair comparison. Materials: Interconnected (CAD-designed, e.g., 3D printed PCL) and conventional (e.g., salt-leached PCL) scaffolds (5x5x2 mm), cell suspension (e.g., hMSCs, 2x10^6 cells/mL), spinner flask or bioreactor, complete growth medium. Procedure:
- Pre-wetting: Place both scaffold types under vacuum in sterile PBS for 30 min to remove trapped air. Transfer to complete medium for 1 hour.
- Scaffold Fixation: Mount each scaffold onto a needle holder within a spinner flask.
- Seeding: Introduce 20 mL of cell suspension into the flask. Stir at 40 rpm for 90 minutes.
- Post-seeding Culture: Transfer scaffolds to a 24-well plate with fresh medium. Culture under static conditions for 24h before assay initiation.
- Efficiency Quantification: Lyse a subset of scaffolds (n=3 per type) at 24h and use a Picogreen dsDNA assay to determine actual cell number versus input.
Protocol B: Assessment of Mass Transport and Cell Infiltration
Objective: To quantify diffusion and cell migration into the scaffold interior. Materials: Scaffolds, fluorescent dextran (70 kDa, FITC-labeled), culture medium, confocal microscope, cryostat. Diffusion Assay (FRAP):
- Equilibrate acellular scaffolds in dextran solution (1 mg/mL) for 24h.
- Using a confocal microscope, photobleach a defined region 500 µm deep into the scaffold.
- Monitor fluorescence recovery over 30 minutes. Calculate effective diffusivity (D/D₀) using standard models. Infiltration Assay:
- Seed scaffolds per Protocol A with GFP-expressing cells.
- At days 3, 7, and 14, fix scaffolds (4% PFA), section vertically (cryostat or vibratome).
- Image sections via confocal microscopy. Measure the distance from the surface where cell density drops to 50% (Infiltration Depth).
Protocol C: Functional Readouts: Metabolism and Gene Expression
Objective: To evaluate the biological impact of scaffold architecture on cell function. Metabolic Activity (AlamarBlue):
- Culture cell-seeded scaffolds in 24-well plates (n=4 per group/time point).
- At each time point, incubate with 10% AlamarBlue reagent in serum-free medium for 3h at 37°C.
- Measure fluorescence (Ex 560/Em 590) of the supernatant. Normalize to day 1 readings. Gene Expression Analysis (qPCR):
- At day 7, homogenize scaffolds (n=4) in TRIzol reagent. Extract total RNA.
- Synthesize cDNA. Perform qPCR for target genes (e.g., VEGFA, COL1A1, RUNX2 for osteogenesis).
- Normalize to housekeeping genes (GAPDH, HPRT1) and calculate relative expression (2^-ΔΔCt) vs. conventional scaffold control.
Visualization of Key Pathways and Workflows
Diagram 1: Scaffold Arch. Impact on Cell Outcomes (76 chars)
Diagram 2: Experimental Workflow for Comparison (73 chars)
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Comparative Scaffold Studies
| Item | Function / Rationale | Example Product/Catalog |
|---|---|---|
| CAD Software & 3D Printer | Designs and fabricates precise interconnected lattice structures (e.g., Gyroid, Diamond). | Autodesk Fusion 360; 3D-Bioplotter (EnvisionTEC). |
| Bioink / Filament | Material for scaffold fabrication. Must be biocompatible and printable. | Medical-grade PCL pellets (Sigma, 704192) or GelMA bioink. |
| Fluorescent Tracer (70 kDa Dextran) | High molecular weight probe to simulate nutrient/protein diffusion without rapid leakage. | FITC-Dextran, 70 kDa (Thermo Fisher, D1822). |
| Live/Dead Viability Assay | Distinguishes live (calcein-AM, green) from dead (ethidium homodimer-1, red) cells in 3D. | Thermo Fisher, L3224. |
| AlamarBlue Cell Viability Reagent | Resazurin-based; non-toxic, allows longitudinal tracking of metabolic activity in the same scaffold. | Thermo Fisher, DAL1100. |
| Total RNA Isolation Reagent for Biomaterials | Efficiently lyses cells within a 3D polymer matrix and isolates intact RNA. | TRIzol Reagent (Thermo Fisher, 15596026). |
| Collagen Type I Antibody | Immunostaining to assess de novo extracellular matrix deposition by cells within scaffolds. | Abcam, ab34710. |
Introduction & Thesis Context The advancement of computer-aided design (CAD) for scaffolds with fully interconnected channel networks is a cornerstone of tissue engineering research. The central thesis posits that predefined, optimized channel architectures directly dictate cellular colonization and subsequent tissue formation. This application note provides standardized protocols and metrics to quantitatively validate this thesis by assessing three critical performance parameters: cell infiltration depth and distribution, spatial uniformity of cell settlement, and resultant metabolic activity. These metrics serve as the essential biological feedback loop for iterative CAD scaffold design.
1.0 Metric 1: Quantifying Cell Infiltration Depth and Distribution
Protocol 1.1: Fluorescent Staining and Confocal Microscopy Analysis for Infiltration Objective: To measure the depth and spatial distribution of viable cells within a 3D scaffold over time. Materials: Cell-seeded scaffold, Calcein AM (viability stain), Phalloidin (actin cytoskeleton), Hoechst 33342 (nuclei), 4% Paraformaldehyde, PBS, Confocal Laser Scanning Microscope (CLSM). Procedure:
- At designated time points (e.g., 1, 7, 14 days), rinse scaffolds 2x with PBS.
- Fix samples with 4% PFA for 30 minutes at room temperature (RT). Rinse 3x with PBS.
- Permeabilize with 0.1% Triton X-100 in PBS for 15 minutes (RT). Rinse.
- Stain with Phalloidin (1:200) and Hoechst (1:1000) in PBS for 60 minutes (RT), protected from light.
- Rinse thoroughly with PBS. For live/dead assays, skip fixation and incubate with Calcein AM (2 µM) for 30-45 minutes.
- Image using a CLSM with Z-stacking capability. Set Z-step interval to 10-20 µm to capture the full scaffold depth.
- Analyze images using Fiji/ImageJ software. Use the "Plot Z-axis Profile" function for fluorescence intensity to generate infiltration profiles.
Data Presentation: Cell Infiltration Metrics Table 1: Quantitative Infiltration Data from CLSM Z-Stacks
| Scaffold Channel Design | Mean Infiltration Depth (µm) Day 7 | Max Infiltration Depth (µm) Day 7 | Gradient Coefficient (R² of Intensity Slope) |
|---|---|---|---|
| Orthogonal Grid (300µm) | 850 ± 120 | 1100 ± 150 | 0.95 ± 0.03 |
| Gyroid (500µm) | 1250 ± 95 | 1550 ± 200 | 0.87 ± 0.05 |
| Radial Spoke | 650 ± 200 | 900 ± 250 | 0.65 ± 0.12 |
| Solid Control (No Channels) | 150 ± 50 | 200 ± 75 | 0.98 ± 0.01 |
2.0 Metric 2: Assessing Spatial Uniformity of Cell Distribution
Protocol 2.1: DNA Quantification Across Scaffold Sections Objective: To obtain a biochemical measure of cell number uniformity across different spatial segments of a scaffold. Materials: Cell-seeded scaffold, Scalpel, PBS, DNA Quantification Kit (e.g., PicoGreen), Microplate Reader, Homogenizer. Procedure:
- Aseptically dissect the cell-seeded scaffold into three equal segments: Top, Middle, and Bottom.
- Homogenize each segment separately in 1 mL of lysis buffer (e.g., with 0.1% SDS).
- Centrifuge homogenates at 10,000xg for 10 minutes to pellet debris.
- Following the PicoGreen kit instructions, mix 100 µL of supernatant with 100 µL of PicoGreen working solution in a black 96-well plate.
- Incubate for 5 minutes, protected from light.
- Measure fluorescence (excitation ~480 nm, emission ~520 nm) using a microplate reader.
- Calculate DNA concentration from a standard curve. Normalize to the wet weight of each scaffold segment.
Data Presentation: Cell Distribution Uniformity Table 2: DNA Content as a Measure of Spatial Cell Distribution (Day 14)
| Scaffold Design | Top Segment (ng DNA/mg) | Middle Segment (ng DNA/mg) | Bottom Segment (ng DNA/mg) | Coefficient of Variation (CV%) Across Segments |
|---|---|---|---|---|
| Orthogonal Grid | 45.2 ± 5.1 | 42.8 ± 4.3 | 40.1 ± 6.0 | 6.5% |
| Gyroid | 52.3 ± 3.8 | 50.1 ± 4.5 | 48.9 ± 5.2 | 3.8% |
| Radial Spoke | 60.1 ± 7.2 | 35.5 ± 6.8 | 22.3 ± 5.5 | 52.1% |
| Solid Control | 28.5 ± 3.2 | 5.1 ± 1.5 | 2.8 ± 0.9 | 108.3% |
3.0 Metric 3: Measuring Metabolic Activity & Viability
Protocol 3.1: Metabolic Activity Assay (AlamarBlue/Resazurin) Objective: To assess the metabolic activity of cells within the scaffold as a proxy for viability and proliferation. Materials: Cell-seeded scaffold, AlamarBlue reagent, Phenol-red free culture medium, Microplate reader, Orbital shaker. Procedure:
- Prepare a 10% (v/v) solution of AlamarBlue reagent in pre-warmed, phenol-red free medium.
- Aspirate culture medium from scaffolds and gently rinse with PBS.
- Add the 10% AlamarBlue solution to completely cover each scaffold.
- Incubate at 37°C for 2-4 hours on an orbital shaker (low speed) to ensure reagent penetration.
- After incubation, transfer 100 µL of the reacted solution from each well to a black or clear 96-well plate.
- Measure fluorescence (Excitation 560 nm, Emission 590 nm) or absorbance (570 nm, reference 600 nm).
- The reduction percentage is calculated per kit instructions. Always include a scaffold-only control in AlamarBlue to account for background.
Data Presentation: Metabolic Activity Over Time Table 3: Metabolic Activity (AlamarBlue % Reduction) Over Culture Period
| Time Point | Gyroid Scaffold | Orthogonal Grid | Acellular Control |
|---|---|---|---|
| Day 1 | 15.2% ± 2.1% | 14.8% ± 1.9% | 1.5% ± 0.5% |
| Day 7 | 68.5% ± 5.3% | 55.7% ± 4.8% | 1.8% ± 0.6% |
| Day 14 | 92.3% ± 3.1% | 78.9% ± 6.2% | 2.1% ± 0.7% |
The Scientist's Toolkit: Research Reagent Solutions Table 4: Essential Materials for Performance Quantification
| Item | Function/Application |
|---|---|
| Calcein AM | Cell-permeant fluorescent dye hydrolyzed by intracellular esterases to indicate viable cells (green). |
| Picogreen dsDNA Assay | Ultra-sensitive fluorescent nucleic acid stain for quantifying low-level DNA, directly correlating to cell number. |
| AlamarBlue (Resazurin) | Cell-permeable redox indicator; reduction by metabolically active cells changes color/fluorescence. |
| Phalloidin (Conjugates) | High-affinity actin filament stain for visualizing the cytoskeleton and cell morphology within scaffolds. |
| Hoechst 33342 | Cell-permeant nuclear counterstain (blue) for identifying total cell distribution. |
| Matrigel or Collagen I | Often used as a hydrogel coating to functionalize synthetic scaffold surfaces for improved cell adhesion. |
| Triton X-100 | Non-ionic detergent used for permeabilizing cell membranes to allow entry of large staining molecules. |
Visualization: Experimental & Analytical Workflows
Title: Cell Infiltration Analysis Protocol
Title: Scaffold Design Biofeedback Loop
This document provides Application Notes and Protocols for advanced functional testing of tissue-engineered scaffolds, framed within a broader thesis on CAD design for scaffolds with fully interconnected channel networks. The efficacy of such computationally designed architectures must be validated through rigorous biomimetic perfusion studies and precise quantification of degradation kinetics. These protocols are designed for researchers, scientists, and drug development professionals to standardize the assessment of scaffold performance under dynamic culture conditions.
Key Research Reagent Solutions & Materials
Table 1: Essential Research Toolkit for Perfusion & Degradation Studies
| Item | Function & Explanation |
|---|---|
| Tri-axial Perfusion Bioreactor | Provides controlled, laminar medium flow through 3D scaffold channels, simulating vascular shear stress. Essential for testing CAD-designed interconnectivity. |
| Poly(D,L-lactide-co-glycolide) (PLGA) Scaffolds | Model biodegradable polymer with tunable degradation rates (via LA:GA ratio). The primary test substrate for degradation kinetics. |
| Fluorescently-Tagged Albumin (e.g., FITC-BSA) | A perfusion tracer molecule. Used to quantify fluid dynamics, distribution efficiency, and confirm channel interconnectivity via fluorescence imaging. |
| Collagenase Type II | Enzyme solution for in vitro accelerated degradation studies. Simulates enzymatic hydrolytic degradation of collagen-based or susceptible polymeric scaffolds. |
| AlamarBlue or PrestoBlue | Cell viability/ metabolic activity resazurin-based assay. For non-destructive, longitudinal monitoring of cell health within perfused scaffolds. |
| Micro-CT Scanner | For non-destructive, high-resolution 3D imaging of scaffold architecture (porosity, channel connectivity) pre- and post-degradation/perfusion. |
| pH-Stat Titration System | Automatically monitors and titrates pH of degradation medium. Precisely quantifies hydrolytic degradation rate by measuring acid release. |
| PCR Primers for Hypoxia/Vascular Markers (e.g., HIF-1α, VEGF) | To assess cellular genetic response to perfusion conditions vs. static culture, validating the biomimetic environment. |
Application Note: Perfusion Bioreactor Protocol for Interconnected Channel Assessment
Objective: To validate the mass transport efficacy and biomimetic shear stress application of a CAD-designed scaffold with interconnected channels using a custom tri-axial perfusion bioreactor system.
Detailed Protocol:
A. Scaffold Preparation & Sterilization
- Fabricate scaffold (e.g., via 3D printing, salt leaching) based on CAD model. Material: PLGA (85:15 LA:GA).
- Measure dry mass (M₀) and dimensions. Image via micro-CT to obtain baseline architecture data.
- Sterilize by immersion in 70% ethanol for 30 minutes, followed by triple rinse in sterile PBS.
- Pre-wet scaffold in culture medium overnight under vacuum to ensure channel infiltration.
B. Bioreactor Setup & Seeding
- Aseptically assemble bioreactor cartridge, ensuring inlet and outlet tubing are securely connected to the scaffold chamber.
- Seed scaffolds with primary human mesenchymal stem cells (hMSCs) at a density of 5 x 10⁵ cells/scaffold using a dynamic seeding method:
- Inject cell suspension into the scaffold chamber.
- Place cartridge on a rocking platform for 2 hours (15 min intervals, flip 180°).
- Transfer cartridge to bioreactor base. Connect to medium reservoir and peristaltic pump.
- Initiate perfusion after a 24-hour static period. Set initial flow rate (Q) to 0.1 mL/min to allow cell adhesion.
C. Perfusion Culture & Monitoring
- Gradually increase flow rate over 72 hours to the target shear stress (τ), calculated using: τ = (4μQ)/(πr³) (for a cylindrical channel approximation, where μ=medium viscosity, r=mean channel radius from CAD). Target τ: 0.5 - 5 mPa (mimicking bone marrow sinusoid shear).
- Maintain culture at 37°C, 5% CO₂. Replace 50% of medium in the reservoir every 48 hours.
- At designated time points (Days 1, 7, 14, 21):
- Viability: Sample effluent medium for lactate dehydrogenase (LDH) assay. Perform AlamarBlue assay on a sacrificial scaffold.
- Perfusion Efficacy: Introduce FITC-BSA (0.1 mg/mL) into the medium reservoir. Collect outlet medium at timed intervals to generate a clearance curve. Image scaffold via confocal microscopy to visualize distribution.
- Cell Response: Harvest RNA from scaffolds for qPCR analysis of osteogenic (Runx2, ALP) and hypoxia (HIF-1α) markers.
D. Data Analysis
- Calculate Distribution Efficiency (DE%) = (Fluorescence intensity at scaffold core / Intensity at periphery) x 100 from confocal slices.
- Correlate DE% with CAD-predicted channel connectivity metric (e.g., Tortuosity Factor).
Diagram 1: Perfusion Bioreactor Experimental Workflow (100 chars)
Application Note: Degradation Kinetics Protocol
Objective: To quantitatively characterize the degradation profile of a CAD-designed porous scaffold, linking mass loss, mechanical decay, and byproduct release to initial architectural parameters.
Detailed Protocol:
A. In Vitro Degradation Study Setup
- Prepare scaffold samples (n=6 per time point, e.g., PLGA). Record initial dry mass (M₀), dimensions, and perform baseline micro-CT/compressive modulus (E₀) testing.
- For Hydrolytic Degradation:
- Immerse each scaffold in 5 mL of sterile PBS (pH 7.4, with 0.02% sodium azide).
- Place in shaking incubator (37°C, 60 rpm).
- For Enzymatic Degradation (Accelerated):
- Immerse in 5 mL of PBS containing 1 U/mL Collagenase Type II (for collagen/ susceptible polymers).
- Include enzyme-free controls.
B. Monitoring & Sampling
- Medium Analysis (Weekly):
- pH Monitoring: Use pH-stat or record pH change. Replace medium with fresh solution to maintain sink conditions.
- Mass Loss: At pre-defined time points (1, 2, 4, 8, 12, 16 weeks), remove one set of samples (n=3). Rinse, lyophilize, and record dry mass (Mt).
- Mechanical Testing: Perform unconfined compression on hydrated samples to determine modulus (Et).
- Morphology: Image via SEM and micro-CT to track pore morphology and channel structure changes.
C. Data Quantification
- Calculate Percent Mass Remaining = (M_t / M₀) x 100.
- Calculate Percent Modulus Remaining = (E_t / E₀) x 100.
- Plot degradation profiles. Model kinetics using empirical models (e.g., first-order exponential decay).
Table 2: Typical Degradation Data for PLGA (85:15) Scaffolds Over 12 Weeks
| Time (Weeks) | Avg. Mass Remaining (%) | Avg. Modulus Remaining (%) | Medium pH | Key Structural Change (Micro-CT) |
|---|---|---|---|---|
| 0 | 100.0 ± 1.5 | 100.0 ± 8.2 | 7.40 | N/A |
| 2 | 98.5 ± 2.1 | 95.3 ± 7.5 | 7.38 | No change |
| 4 | 96.8 ± 1.8 | 88.7 ± 9.1 | 7.35 | No change |
| 8 | 85.4 ± 3.5 | 62.1 ± 10.4 | 7.22 | Initial pore wall thinning |
| 12 | 60.2 ± 5.7 | 30.5 ± 12.8 | 7.05 | Loss of small struts, channel merging |
Diagram 2: Hydrolytic Degradation Pathway & Effects (99 chars)
Integrated Test: Perfusion-Driven Degradation
Objective: To study the coupled effect of dynamic fluid flow on scaffold degradation kinetics, mimicking a more physiologically relevant environment.
Protocol Summary:
- Place sterile scaffolds in perfusion bioreactors. Use degradation medium (PBS + antibiotics) as perfusate.
- Apply a low, constant shear stress (0.2 mPa).
- At intervals, sample outlet medium for pH and lactic acid quantification (HPLC).
- Compare mass loss and micro-CT structural changes against static degradation controls (from Section 4).
- Expected Outcome: Perfusion may accelerate degradation due to enhanced convective removal of acidic oligomers, reducing autocatalytic effects in the scaffold core. This directly tests how channel interconnectivity influences the degradation homogeneity.
This document provides a structured framework for the comparative benchmarking of novel CAD-designed scaffolds against established state-of-the-art (SOTA) commercial and research scaffolds, within the context of a thesis focused on designing scaffolds with fully interconnected channel networks.
1.0 Core Quantitative Comparison Table Table 1. Benchmarking Metrics for Scaffold Evaluation
| Metric Category | Specific Parameter | Novel CAD Design (Typical Target) | Commercial SOTA (e.g., NuVasive Modulus, Ossiform BONITmatrix) | Research SOTA (e.g., Triply Periodic Minimal Surface (TPMS)) |
|---|---|---|---|---|
| Architectural | Porosity (%) | 70-85% (Designed) | 55-75% (Stochastic) | 70-90% (Designed) |
| Pore Size (µm) | 300-600 (Isotropic/Anisotropic) | 100-800 (Stochastic) | 200-1000 (Precise) | |
| Interconnectivity | Fully Interconnected Network (Designed) | High (Stochastic) | Fully Interconnected (Designed) | |
| Mechanical | Compressive Modulus (MPa) | 0.5-3.0 (Soft Tissue); 50-500 (Bone) | 0.1-2.0 (Collagen); 100-2000 (HA/TCP) | Tunable across range |
| Permeability (m²) | 1e-10 - 1e-8 (Designed) | 1e-11 - 1e-9 | 1e-10 - 1e-8 | |
| Biological Performance | Cell Seeding Efficiency (%) | >90% (Channel-Enhanced) | 60-80% | 70-95% |
| Metabolic Activity (Day 7) | 150-200% vs. Control | 100-130% vs. Control | 120-180% vs. Control | |
| Mineralization (Bone, Day 21) µg/mL | ~300 (Calcium Content) | ~150-250 | ~250-350 |
2.0 Detailed Experimental Protocols
Protocol 2.1: Architectural and Permeability Benchmarking Objective: Quantify pore architecture and fluid transport against SOTA controls. Materials: Micro-CT scanner (e.g., Bruker SkyScan), ImageJ with BoneJ plugin, Computational Fluid Dynamics (CFD) software (e.g., ANSYS Fluent). Procedure:
- Imaging: Scan all scaffold groups (n=5/group) at isotropic voxel size ≤10µm.
- Architectural Analysis: Reconstruct 3D models. Use BoneJ to calculate porosity, pore size distribution, and strut thickness.
- Interconnectivity Verification: Apply a 3D connectivity analysis. Label all connected pores; a fully interconnected design yields one single label.
- Permeability Simulation: Import segmented 3D model into CFD. Apply a pressure gradient (∆P = 10 Pa) across the scaffold model in a simulated fluid (viscosity = 0.001 Pa·s). Solve Stokes flow to calculate Darcy permeability (k).
Protocol 2.2: In Vitro Biological Performance Benchmarking Objective: Compare cell migration, proliferation, and differentiation. Materials: Human Mesenchymal Stem Cells (hMSCs), osteogenic media, AlamarBlue assay, Live/Dead staining kit, microplate reader. Procedure:
- Sterilization & Pre-wetting: Ethanol (70%, 30 min), PBS rinse, culture media incubation (2 hrs).
- Dynamic Seeding: Seed hMSCs (50,000 cells/scaffold) in a spinner flask (25 rpm, 2 hrs). Calculate seeding efficiency: (1 - cells in supernatant/total cells) * 100.
- Culture: Maintain in osteogenic media. Refresh every 3 days.
- Metabolic Activity: At days 1, 4, 7, incubate with AlamarBlue (10% v/v, 2 hrs). Measure fluorescence (Ex560/Em590).
- Live/Dead & Imaging: At day 7, stain with calcein-AM/ethidium homodimer-1. Image via confocal microscopy to visualize cell penetration depth through interconnected channels.
3.0 Visualization of Key Processes
Title: Benchmarking Experimental Workflow
Title: Scaffold Architecture Induces Osteogenesis
4.0 The Scientist's Toolkit: Key Research Reagent Solutions
Table 2. Essential Materials for Scaffold Benchmarking
| Item Name | Supplier Examples | Function in Benchmarking |
|---|---|---|
| Human Mesenchymal Stem Cells (hMSCs) | Lonza, Thermo Fisher | Primary cell model for evaluating osteogenic response. |
| Osteogenic Differentiation Media | MilliporeSigma, STEMCELL Tech | Induces bone formation; standardizes differentiation potential tests. |
| AlamarBlue Cell Viability Reagent | Thermo Fisher, Bio-Rad | Fluorescent redox indicator for non-destructive metabolic activity tracking. |
| Calcein-AM / EthD-1 Live/Dead Kit | Thermo Fisher | Simultaneously stains live (green) and dead (red) cells for viability/penetration. |
| Micro-CT Calibration Phantoms | Bruker, Scanco | Ensures accurate quantification of architectural parameters (porosity, thickness). |
| Image Analysis Software (BoneJ/CTAn) | Open Source, Bruker | Essential plugin for robust, reproducible 3D morphometric analysis from micro-CT data. |
Conclusion
The CAD-driven design of scaffolds with fully interconnected channel networks represents a paradigm shift in tissue engineering and regenerative medicine. By integrating foundational biological principles, advanced methodological workflows, systematic troubleshooting, and rigorous validation, researchers can now engineer biomimetic architectures that were previously unattainable. The key takeaway is that interconnectivity is not merely a structural feature but a functional prerequisite for sustaining life within engineered constructs. Future directions will focus on the integration of machine learning for automated topology optimization, the development of multi-material CAD strategies to create heterogeneous channel microenvironments, and the direct linkage of patient imaging data to scaffold design for truly personalized implants. As these technologies converge, they promise to accelerate the translation of lab-designed scaffolds into clinically viable solutions, fundamentally advancing drug development pipelines and regenerative therapies.